Program: 17. Neuroblastoma #4.
Submit a comment on this gene expression program’s interpretation: CLICK
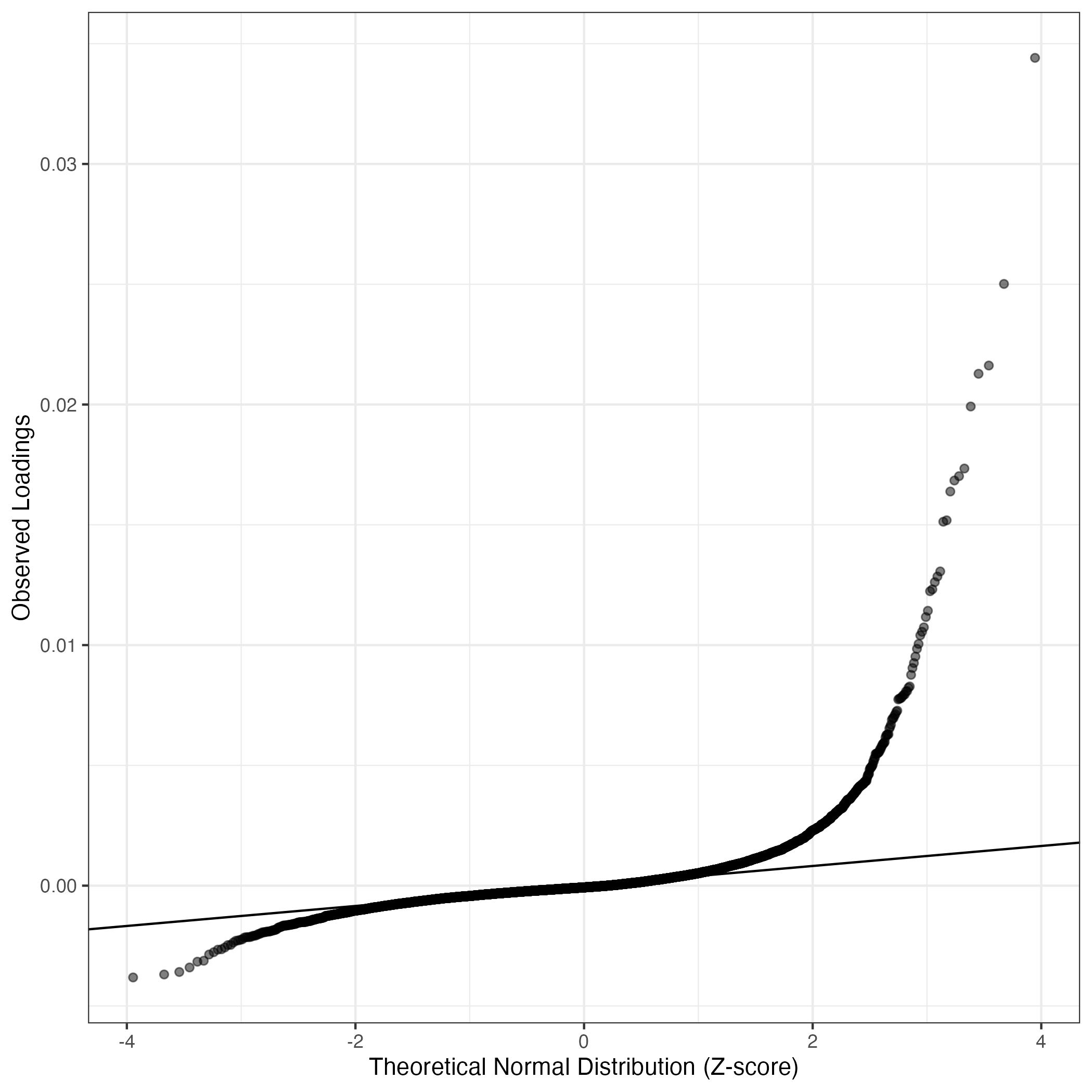
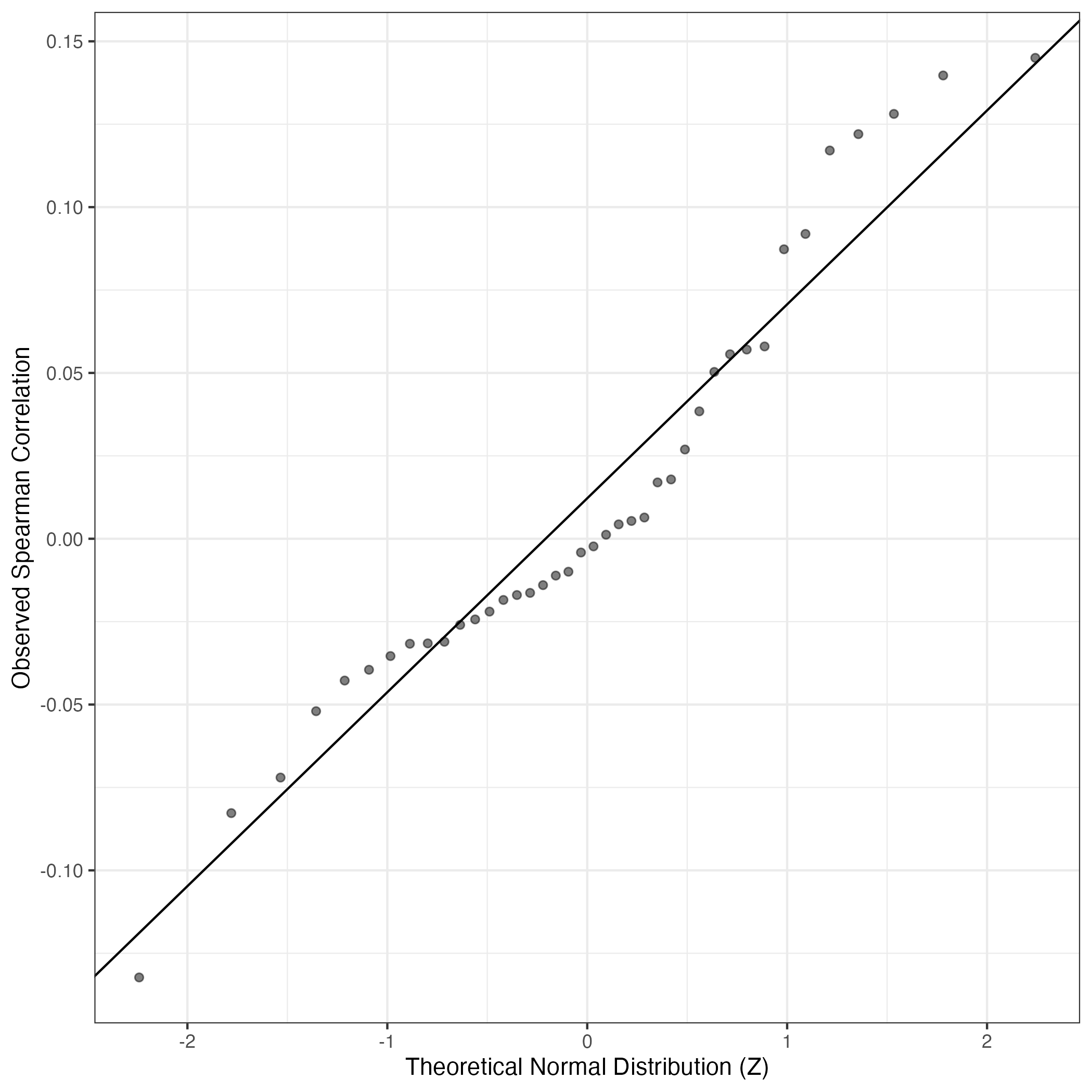
QQ-plot of gene loadings, averaged over both independent splits of the data
This plot highlights the relative contribution of each gene to the GEP
Top genes driving this program.
Note: Decartes website is buggy, try refreshing. Also, Decartes fetal adrenal data have been collected at specific time points (89-122 days), all possible cell types of interest may not be represented, do not overinterpret.
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
| Gene | Loading | Gene.Name | GTEx | DepMap | Descartes | Mean.Counts | Mean.Tpm | |
|---|---|---|---|---|---|---|---|---|
| 1 | SSTR2 | 0.0344108 | somatostatin receptor 2 | GTEx | DepMap | Descartes | 4.98 | 270.38 |
| 2 | SIX1 | 0.0250132 | SIX homeobox 1 | GTEx | DepMap | Descartes | 0.65 | 66.49 |
| 3 | CYP26A1 | 0.0216206 | cytochrome P450 family 26 subfamily A member 1 | GTEx | DepMap | Descartes | 0.56 | 105.51 |
| 4 | PPP1R14A | 0.0212789 | protein phosphatase 1 regulatory inhibitor subunit 14A | GTEx | DepMap | Descartes | 2.25 | 993.31 |
| 5 | LDB3 | 0.0199186 | LIM domain binding 3 | GTEx | DepMap | Descartes | 0.38 | 31.65 |
| 6 | DACH1 | 0.0173399 | dachshund family transcription factor 1 | GTEx | DepMap | Descartes | 1.00 | 78.52 |
| 7 | CHRNA1 | 0.0170226 | cholinergic receptor nicotinic alpha 1 subunit | GTEx | DepMap | Descartes | 0.30 | 48.89 |
| 8 | EYA2 | 0.0168411 | EYA transcriptional coactivator and phosphatase 2 | GTEx | DepMap | Descartes | 0.31 | 50.45 |
| 9 | GKAP1 | 0.0163840 | G kinase anchoring protein 1 | GTEx | DepMap | Descartes | 2.03 | 482.60 |
| 10 | ASB4 | 0.0151877 | ankyrin repeat and SOCS box containing 4 | GTEx | DepMap | Descartes | 0.21 | 30.55 |
| 11 | RAPSN | 0.0151285 | receptor associated protein of the synapse | GTEx | DepMap | Descartes | 0.20 | 54.32 |
| 12 | HES6 | 0.0130626 | hes family bHLH transcription factor 6 | GTEx | DepMap | Descartes | 1.78 | 502.20 |
| 13 | PAPSS1 | 0.0128511 | 3’-phosphoadenosine 5’-phosphosulfate synthase 1 | GTEx | DepMap | Descartes | 1.56 | 248.71 |
| 14 | THSD7B | 0.0126168 | thrombospondin type 1 domain containing 7B | GTEx | DepMap | Descartes | 0.29 | 20.21 |
| 15 | CERKL | 0.0123165 | ceramide kinase like | GTEx | DepMap | Descartes | 0.19 | 25.68 |
| 16 | NEUROD2 | 0.0122400 | neuronal differentiation 2 | GTEx | DepMap | Descartes | 0.13 | 11.45 |
| 17 | ST18 | 0.0114264 | ST18 C2H2C-type zinc finger transcription factor | GTEx | DepMap | Descartes | 0.14 | 9.51 |
| 18 | FHIT | 0.0111671 | fragile histidine triad diadenosine triphosphatase | GTEx | DepMap | Descartes | 1.02 | 130.23 |
| 19 | ASS1 | 0.0107363 | argininosuccinate synthase 1 | GTEx | DepMap | Descartes | 0.34 | 64.87 |
| 20 | DLK1 | 0.0105537 | delta like non-canonical Notch ligand 1 | GTEx | DepMap | Descartes | 3.96 | 286.85 |
| 21 | SMAD9 | 0.0104037 | SMAD family member 9 | GTEx | DepMap | Descartes | 0.68 | 45.97 |
| 22 | RAI14 | 0.0100483 | retinoic acid induced 14 | GTEx | DepMap | Descartes | 0.72 | 45.97 |
| 23 | LSAMP | 0.0098449 | limbic system associated membrane protein | GTEx | DepMap | Descartes | 0.90 | 40.53 |
| 24 | PEX5L | 0.0095229 | peroxisomal biogenesis factor 5 like | GTEx | DepMap | Descartes | 0.32 | 14.71 |
| 25 | ANO3 | 0.0092548 | anoctamin 3 | GTEx | DepMap | Descartes | 0.18 | 12.38 |
| 26 | MYBPHL | 0.0090470 | myosin binding protein H like | GTEx | DepMap | Descartes | 0.05 | 17.64 |
| 27 | SVIP | 0.0087631 | small VCP interacting protein | GTEx | DepMap | Descartes | 1.40 | 106.22 |
| 28 | IGFBP2 | 0.0082801 | insulin like growth factor binding protein 2 | GTEx | DepMap | Descartes | 3.84 | 312.04 |
| 29 | NKAIN4 | 0.0082292 | sodium/potassium transporting ATPase interacting 4 | GTEx | DepMap | Descartes | 0.49 | 121.93 |
| 30 | AFAP1 | 0.0080883 | actin filament associated protein 1 | GTEx | DepMap | Descartes | 0.75 | 37.49 |
| 31 | RBFOX3 | 0.0080744 | RNA binding fox-1 homolog 3 | GTEx | DepMap | Descartes | 0.24 | 31.25 |
| 32 | GNA14 | 0.0079474 | G protein subunit alpha 14 | GTEx | DepMap | Descartes | 0.12 | 21.40 |
| 33 | GRIN2D | 0.0079274 | glutamate ionotropic receptor NMDA type subunit 2D | GTEx | DepMap | Descartes | 0.12 | 6.94 |
| 34 | GFRA1 | 0.0078773 | GDNF family receptor alpha 1 | GTEx | DepMap | Descartes | 0.26 | 11.91 |
| 35 | CLVS1 | 0.0078117 | clavesin 1 | GTEx | DepMap | Descartes | 0.34 | 38.55 |
| 36 | SATB2 | 0.0077829 | SATB homeobox 2 | GTEx | DepMap | Descartes | 0.08 | 5.60 |
| 37 | GIPR | 0.0077754 | gastric inhibitory polypeptide receptor | GTEx | DepMap | Descartes | 0.07 | 10.24 |
| 38 | DLL3 | 0.0077411 | delta like canonical Notch ligand 3 | GTEx | DepMap | Descartes | 1.06 | 189.59 |
| 39 | PCDH17 | 0.0072739 | protocadherin 17 | GTEx | DepMap | Descartes | 1.00 | 46.29 |
| 40 | DDX1 | 0.0072310 | DEAD-box helicase 1 | GTEx | DepMap | Descartes | 77.91 | 8217.05 |
| 41 | ROBO1 | 0.0071305 | roundabout guidance receptor 1 | GTEx | DepMap | Descartes | 0.51 | 24.25 |
| 42 | NAV2 | 0.0070679 | neuron navigator 2 | GTEx | DepMap | Descartes | 0.35 | 11.28 |
| 43 | NNAT | 0.0069634 | neuronatin | GTEx | DepMap | Descartes | 3.03 | 975.43 |
| 44 | FAM78B | 0.0069573 | family with sequence similarity 78 member B | GTEx | DepMap | Descartes | 0.46 | 35.64 |
| 45 | FZD7 | 0.0068924 | frizzled class receptor 7 | GTEx | DepMap | Descartes | 0.22 | 19.51 |
| 46 | PSMB5 | 0.0066929 | proteasome 20S subunit beta 5 | GTEx | DepMap | Descartes | 4.19 | 1180.09 |
| 47 | LDHA | 0.0065988 | lactate dehydrogenase A | GTEx | DepMap | Descartes | 6.31 | 864.39 |
| 48 | ENC1 | 0.0065283 | ectodermal-neural cortex 1 | GTEx | DepMap | Descartes | 0.49 | 33.15 |
| 49 | KCNJ6 | 0.0062943 | potassium inwardly rectifying channel subfamily J member 6 | GTEx | DepMap | Descartes | 0.11 | NA |
| 50 | STARD4 | 0.0062812 | StAR related lipid transfer domain containing 4 | GTEx | DepMap | Descartes | 0.39 | 34.18 |
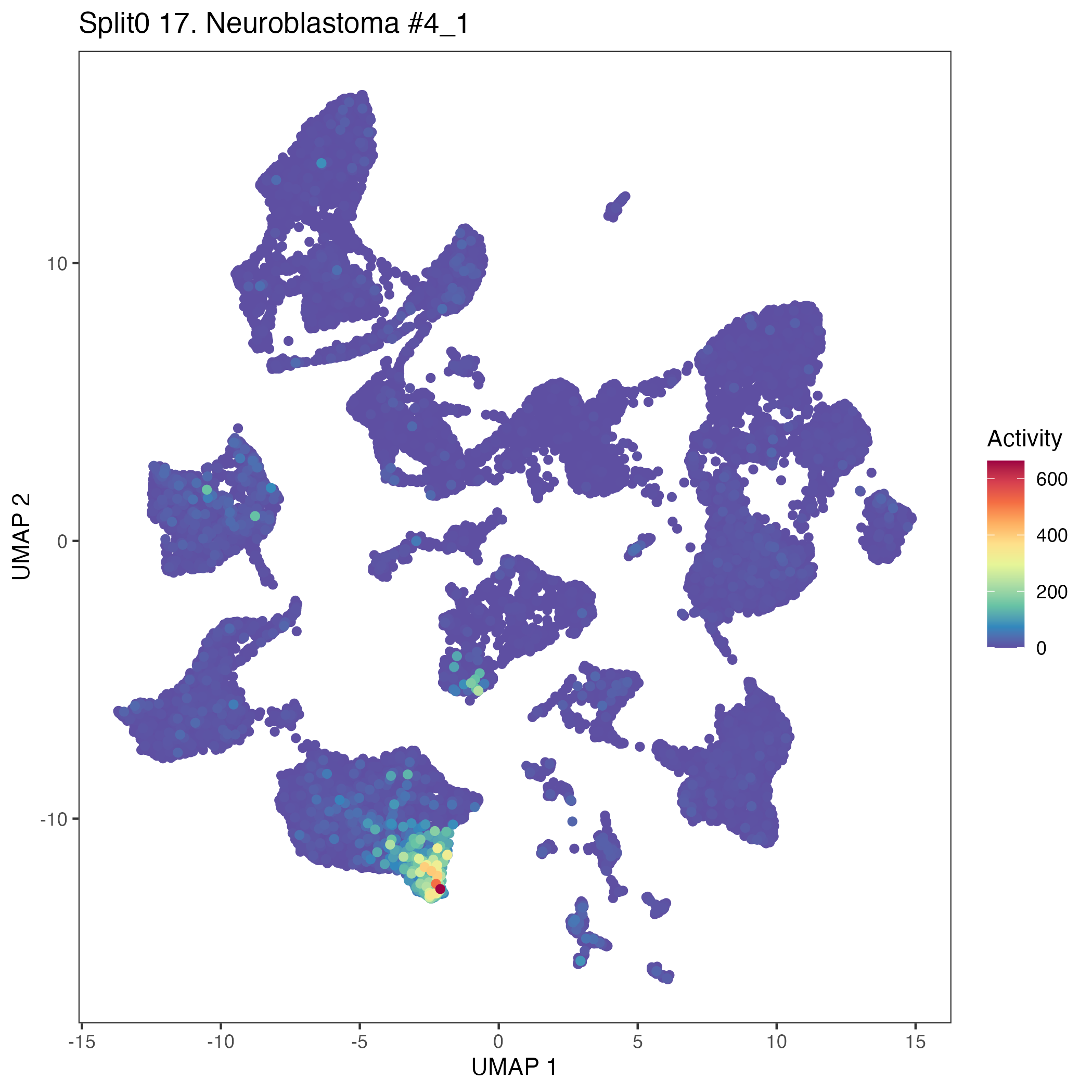
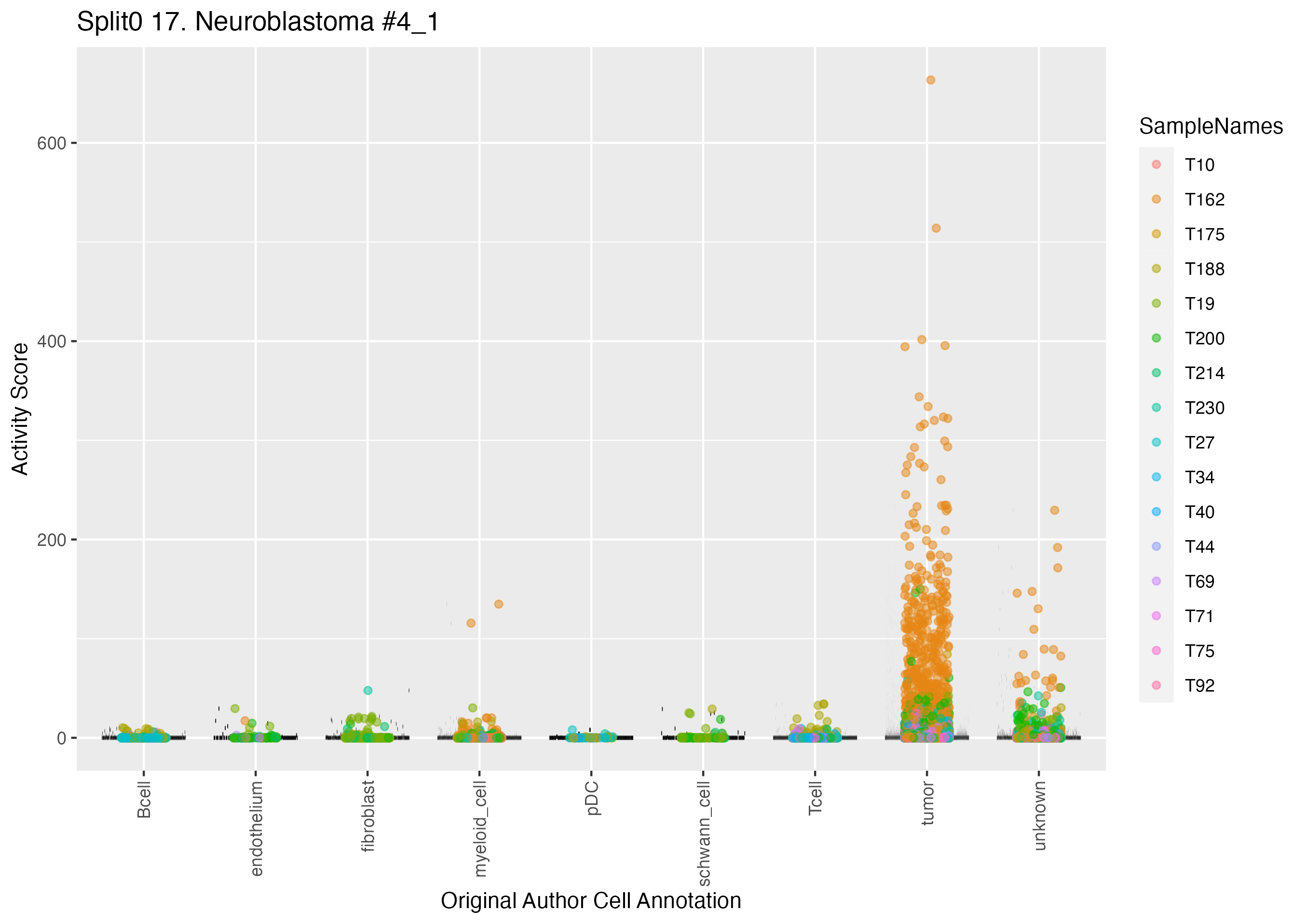
UMAP plots showing activity of gene expression program identified in GEP 17. Neuroblastoma #4:
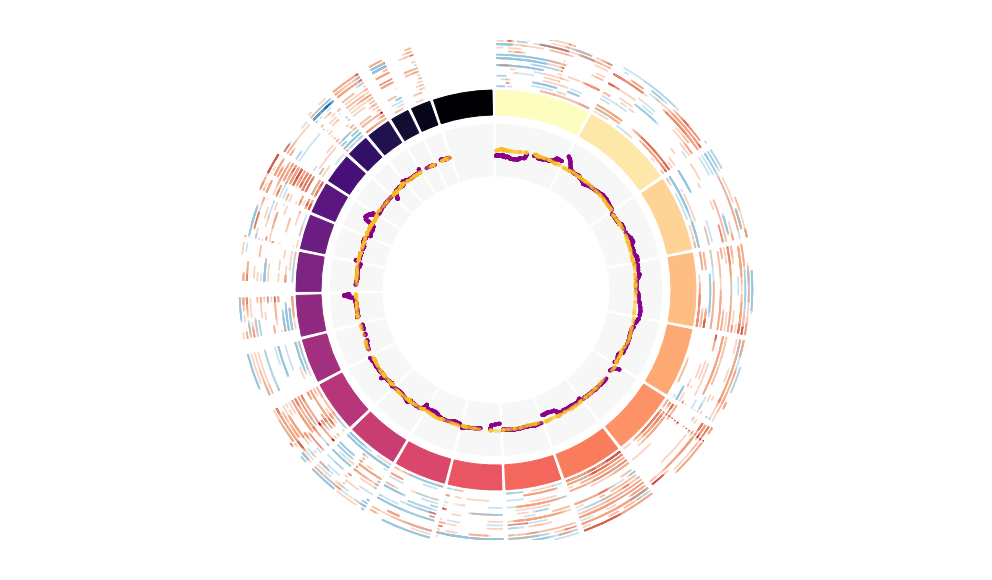
CNV Data procured from inferCNV.
Outer tracks are putative CNV regions (gains = red, losses = blue) for each patient
Inner track is expression data representing:
The top cells expressing this GEP (purple)
Random cells (n =50) from the reference set used in inferCNV (orange)
Gene set Enrichments for this program, caculated from top 50 genes
mSigDB Cell Types Gene Set:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| ZHONG_PFC_C2_SOX5_BCL11B_POS_EXCITATORY_NEURON | 1.81e-04 | 31.85 | 5.95 | 1.35e-02 | 1.21e-01 | 3ST18, LSAMP, LDHA |
28 |
| MANNO_MIDBRAIN_NEUROTYPES_HNBM | 1.31e-08 | 12.42 | 5.67 | 8.80e-06 | 8.80e-06 | 11SSTR2, ST18, PEX5L, SVIP, AFAP1, RBFOX3, GFRA1, CLVS1, DLL3, ROBO1, NNAT |
295 |
| DESCARTES_FETAL_INTESTINE_CHROMAFFIN_CELLS | 7.64e-05 | 12.97 | 3.94 | 1.03e-02 | 5.13e-02 | 5SSTR2, CERKL, ST18, PEX5L, GIPR |
112 |
| DESCARTES_FETAL_PANCREAS_ISLET_ENDOCRINE_CELLS | 4.77e-05 | 10.52 | 3.61 | 9.64e-03 | 3.20e-02 | 6EYA2, CERKL, NEUROD2, ST18, GIPR, KCNJ6 |
168 |
| FAN_EMBRYONIC_CTX_BIG_GROUPS_EXCITATORY_NEURON | 8.44e-04 | 18.11 | 3.48 | 5.66e-02 | 5.66e-01 | 3NEUROD2, SATB2, ENC1 |
47 |
| HU_FETAL_RETINA_RGC | 4.15e-05 | 6.32 | 2.69 | 9.64e-03 | 2.79e-02 | 9DACH1, PAPSS1, SMAD9, NKAIN4, AFAP1, RBFOX3, DLL3, NAV2, LDHA |
443 |
| DESCARTES_FETAL_CEREBRUM_EXCITATORY_NEURONS | 4.05e-03 | 23.65 | 2.62 | 2.09e-01 | 1.00e+00 | 2NEUROD2, SATB2 |
24 |
| ZHONG_PFC_C7_ORG_UNDERGOING_NEURONAL_DIFFERENTIATION | 2.35e-03 | 12.46 | 2.42 | 1.31e-01 | 1.00e+00 | 3NEUROD2, SATB2, ENC1 |
67 |
| MANNO_MIDBRAIN_NEUROTYPES_HDA1 | 5.75e-05 | 5.45 | 2.42 | 9.64e-03 | 3.86e-02 | 10SSTR2, ASB4, DLK1, LSAMP, GFRA1, CLVS1, ROBO1, NNAT, ENC1, KCNJ6 |
584 |
| MANNO_MIDBRAIN_NEUROTYPES_HDA | 1.11e-04 | 5.52 | 2.35 | 1.03e-02 | 7.48e-02 | 9SSTR2, THSD7B, DLK1, LSAMP, GFRA1, CLVS1, ROBO1, NNAT, ENC1 |
506 |
| MANNO_MIDBRAIN_NEUROTYPES_HDA2 | 1.23e-04 | 5.45 | 2.31 | 1.03e-02 | 8.27e-02 | 9SSTR2, ASB4, DLK1, PEX5L, GFRA1, PCDH17, ROBO1, NNAT, KCNJ6 |
513 |
| HAY_BONE_MARROW_STROMAL | 9.96e-05 | 4.68 | 2.15 | 1.03e-02 | 6.68e-02 | 11SSTR2, SIX1, THSD7B, ASS1, SMAD9, RAI14, IGFBP2, GFRA1, SATB2, NAV2, FZD7 |
765 |
| DESCARTES_FETAL_LUNG_NEUROENDOCRINE_CELLS | 5.21e-03 | 9.27 | 1.81 | 2.50e-01 | 1.00e+00 | 3HES6, DLL3, KCNJ6 |
89 |
| MANNO_MIDBRAIN_NEUROTYPES_HNBGABA | 1.12e-03 | 3.96 | 1.68 | 6.81e-02 | 7.50e-01 | 9THSD7B, LSAMP, PEX5L, RBFOX3, CLVS1, PCDH17, ROBO1, NNAT, ENC1 |
703 |
| DESCARTES_FETAL_HEART_SATB2_LRRC7_POSITIVE_CELLS | 1.07e-02 | 7.06 | 1.39 | 4.50e-01 | 1.00e+00 | 3HES6, NEUROD2, SATB2 |
116 |
| DESCARTES_FETAL_STOMACH_ENS_GLIA | 1.55e-02 | 11.31 | 1.29 | 5.11e-01 | 1.00e+00 | 2NKAIN4, FAM78B |
48 |
| MANNO_MIDBRAIN_NEUROTYPES_HGABA | 6.01e-03 | 2.86 | 1.27 | 2.69e-01 | 1.00e+00 | 10SSTR2, LSAMP, PEX5L, RBFOX3, GFRA1, CLVS1, PCDH17, ROBO1, NNAT, ENC1 |
1105 |
| MANNO_MIDBRAIN_NEUROTYPES_HNPROG | 1.22e-02 | 4.83 | 1.25 | 4.80e-01 | 1.00e+00 | 4HES6, ST18, CLVS1, DLL3 |
229 |
| DURANTE_ADULT_OLFACTORY_NEUROEPITHELIUM_OLFACTORY_MICROVILLAR_CELLS | 1.87e-02 | 10.20 | 1.17 | 5.24e-01 | 1.00e+00 | 2HES6, ENC1 |
53 |
| DESCARTES_FETAL_THYMUS_THYMIC_EPITHELIAL_CELLS | 1.52e-02 | 4.51 | 1.17 | 5.11e-01 | 1.00e+00 | 4EYA2, THSD7B, NKAIN4, CLVS1 |
245 |
Dowload full table
mSigDB Hallmark Gene Sets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| HALLMARK_MYOGENESIS | 4.35e-02 | 4.05 | 0.80 | 1.00e+00 | 1.00e+00 | 3LDB3, CHRNA1, NAV2 |
200 |
| HALLMARK_MTORC1_SIGNALING | 4.35e-02 | 4.05 | 0.80 | 1.00e+00 | 1.00e+00 | 3PSMB5, LDHA, STARD4 |
200 |
| HALLMARK_ESTROGEN_RESPONSE_EARLY | 1.85e-01 | 2.63 | 0.31 | 1.00e+00 | 1.00e+00 | 2GFRA1, NAV2 |
200 |
| HALLMARK_XENOBIOTIC_METABOLISM | 1.85e-01 | 2.63 | 0.31 | 1.00e+00 | 1.00e+00 | 2CYP26A1, HES6 |
200 |
| HALLMARK_NOTCH_SIGNALING | 1.20e-01 | 8.23 | 0.20 | 1.00e+00 | 1.00e+00 | 1FZD7 |
32 |
| HALLMARK_CHOLESTEROL_HOMEOSTASIS | 2.55e-01 | 3.49 | 0.09 | 1.00e+00 | 1.00e+00 | 1STARD4 |
74 |
| HALLMARK_PI3K_AKT_MTOR_SIGNALING | 3.41e-01 | 2.45 | 0.06 | 1.00e+00 | 1.00e+00 | 1GNA14 |
105 |
| HALLMARK_SPERMATOGENESIS | 4.15e-01 | 1.90 | 0.05 | 1.00e+00 | 1.00e+00 | 1CLVS1 |
135 |
| HALLMARK_FATTY_ACID_METABOLISM | 4.66e-01 | 1.62 | 0.04 | 1.00e+00 | 1.00e+00 | 1LDHA |
158 |
| HALLMARK_UV_RESPONSE_UP | 4.66e-01 | 1.62 | 0.04 | 1.00e+00 | 1.00e+00 | 1IGFBP2 |
158 |
| HALLMARK_HYPOXIA | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1LDHA |
200 |
| HALLMARK_ESTROGEN_RESPONSE_LATE | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1ASS1 |
200 |
| HALLMARK_MYC_TARGETS_V1 | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1LDHA |
200 |
| HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1IGFBP2 |
200 |
| HALLMARK_OXIDATIVE_PHOSPHORYLATION | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1LDHA |
200 |
| HALLMARK_GLYCOLYSIS | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1LDHA |
200 |
| HALLMARK_KRAS_SIGNALING_DN | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1IGFBP2 |
200 |
| HALLMARK_TNFA_SIGNALING_VIA_NFKB | 1.00e+00 | 0.00 | 0.00 | 1.00e+00 | 1.00e+00 | 0 |
200 |
| HALLMARK_MITOTIC_SPINDLE | 1.00e+00 | 0.00 | 0.00 | 1.00e+00 | 1.00e+00 | 0 |
199 |
| HALLMARK_WNT_BETA_CATENIN_SIGNALING | 1.00e+00 | 0.00 | 0.00 | 1.00e+00 | 1.00e+00 | 0 |
42 |
Dowload full table
KEGG Pathways:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| KEGG_NEUROACTIVE_LIGAND_RECEPTOR_INTERACTION | 2.13e-02 | 4.06 | 1.05 | 1.00e+00 | 1.00e+00 | 4SSTR2, CHRNA1, GRIN2D, GIPR |
272 |
| KEGG_SULFUR_METABOLISM | 5.05e-02 | 21.22 | 0.49 | 1.00e+00 | 1.00e+00 | 1PAPSS1 |
13 |
| KEGG_PURINE_METABOLISM | 1.29e-01 | 3.32 | 0.39 | 1.00e+00 | 1.00e+00 | 2PAPSS1, FHIT |
159 |
| KEGG_CALCIUM_SIGNALING_PATHWAY | 1.55e-01 | 2.96 | 0.35 | 1.00e+00 | 1.00e+00 | 2GNA14, GRIN2D |
178 |
| KEGG_SELENOAMINO_ACID_METABOLISM | 9.85e-02 | 10.19 | 0.24 | 1.00e+00 | 1.00e+00 | 1PAPSS1 |
26 |
| KEGG_ALANINE_ASPARTATE_AND_GLUTAMATE_METABOLISM | 1.20e-01 | 8.23 | 0.20 | 1.00e+00 | 1.00e+00 | 1ASS1 |
32 |
| KEGG_PROPANOATE_METABOLISM | 1.23e-01 | 7.97 | 0.19 | 1.00e+00 | 1.00e+00 | 1LDHA |
33 |
| KEGG_CYSTEINE_AND_METHIONINE_METABOLISM | 1.27e-01 | 7.73 | 0.19 | 1.00e+00 | 1.00e+00 | 1LDHA |
34 |
| KEGG_PYRUVATE_METABOLISM | 1.47e-01 | 6.54 | 0.16 | 1.00e+00 | 1.00e+00 | 1LDHA |
40 |
| KEGG_PROTEASOME | 1.67e-01 | 5.67 | 0.14 | 1.00e+00 | 1.00e+00 | 1PSMB5 |
46 |
| KEGG_NOTCH_SIGNALING_PATHWAY | 1.71e-01 | 5.54 | 0.13 | 1.00e+00 | 1.00e+00 | 1DLL3 |
47 |
| KEGG_AMYOTROPHIC_LATERAL_SCLEROSIS_ALS | 1.90e-01 | 4.91 | 0.12 | 1.00e+00 | 1.00e+00 | 1GRIN2D |
53 |
| KEGG_ARGININE_AND_PROLINE_METABOLISM | 1.94e-01 | 4.81 | 0.12 | 1.00e+00 | 1.00e+00 | 1ASS1 |
54 |
| KEGG_NON_SMALL_CELL_LUNG_CANCER | 1.94e-01 | 4.81 | 0.12 | 1.00e+00 | 1.00e+00 | 1FHIT |
54 |
| KEGG_BASAL_CELL_CARCINOMA | 1.97e-01 | 4.72 | 0.12 | 1.00e+00 | 1.00e+00 | 1FZD7 |
55 |
| KEGG_GLYCOLYSIS_GLUCONEOGENESIS | 2.19e-01 | 4.18 | 0.10 | 1.00e+00 | 1.00e+00 | 1LDHA |
62 |
| KEGG_RETINOL_METABOLISM | 2.25e-01 | 4.05 | 0.10 | 1.00e+00 | 1.00e+00 | 1CYP26A1 |
64 |
| KEGG_LONG_TERM_POTENTIATION | 2.43e-01 | 3.70 | 0.09 | 1.00e+00 | 1.00e+00 | 1GRIN2D |
70 |
| KEGG_SMALL_CELL_LUNG_CANCER | 2.84e-01 | 3.07 | 0.08 | 1.00e+00 | 1.00e+00 | 1FHIT |
84 |
| KEGG_TGF_BETA_SIGNALING_PATHWAY | 2.90e-01 | 3.00 | 0.07 | 1.00e+00 | 1.00e+00 | 1SMAD9 |
86 |
Dowload full table
CHR Positional Gene Sets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| chr11p14 | 2.07e-02 | 9.64 | 1.11 | 1.00e+00 | 1.00e+00 | 2ANO3, SVIP |
56 |
| chr2q31 | 1.40e-01 | 3.16 | 0.37 | 1.00e+00 | 1.00e+00 | 2CHRNA1, CERKL |
167 |
| chr9q21 | 1.68e-01 | 2.80 | 0.33 | 1.00e+00 | 1.00e+00 | 2GKAP1, GNA14 |
188 |
| chr10q23 | 1.71e-01 | 2.77 | 0.32 | 1.00e+00 | 1.00e+00 | 2CYP26A1, LDB3 |
190 |
| chr2q33 | 1.78e-01 | 2.70 | 0.32 | 1.00e+00 | 1.00e+00 | 2SATB2, FZD7 |
195 |
| chr19q13 | 1.00e+00 | 0.94 | 0.24 | 1.00e+00 | 1.00e+00 | 4PPP1R14A, GRIN2D, GIPR, DLL3 |
1165 |
| chr17q25 | 3.22e-01 | 1.77 | 0.21 | 1.00e+00 | 1.00e+00 | 2SSTR2, RBFOX3 |
297 |
| chr20q13 | 6.68e-01 | 1.31 | 0.15 | 1.00e+00 | 1.00e+00 | 2EYA2, NKAIN4 |
400 |
| chr5q22 | 1.97e-01 | 4.72 | 0.12 | 1.00e+00 | 1.00e+00 | 1STARD4 |
55 |
| chr3p12 | 2.31e-01 | 3.92 | 0.10 | 1.00e+00 | 1.00e+00 | 1ROBO1 |
66 |
| chr2q22 | 2.37e-01 | 3.81 | 0.09 | 1.00e+00 | 1.00e+00 | 1THSD7B |
68 |
| chr8q11 | 2.43e-01 | 3.70 | 0.09 | 1.00e+00 | 1.00e+00 | 1ST18 |
70 |
| chr2p24 | 2.55e-01 | 3.49 | 0.09 | 1.00e+00 | 1.00e+00 | 1DDX1 |
74 |
| chr13q13 | 2.67e-01 | 3.31 | 0.08 | 1.00e+00 | 1.00e+00 | 1SMAD9 |
78 |
| chr4q25 | 2.93e-01 | 2.97 | 0.07 | 1.00e+00 | 1.00e+00 | 1PAPSS1 |
87 |
| chr8q12 | 2.95e-01 | 2.93 | 0.07 | 1.00e+00 | 1.00e+00 | 1CLVS1 |
88 |
| chr3p14 | 3.84e-01 | 2.11 | 0.05 | 1.00e+00 | 1.00e+00 | 1FHIT |
122 |
| chr1q24 | 3.87e-01 | 2.09 | 0.05 | 1.00e+00 | 1.00e+00 | 1FAM78B |
123 |
| chr14q23 | 3.89e-01 | 2.07 | 0.05 | 1.00e+00 | 1.00e+00 | 1SIX1 |
124 |
| chr10q25 | 3.94e-01 | 2.04 | 0.05 | 1.00e+00 | 1.00e+00 | 1GFRA1 |
126 |
Dowload full table
Transcription Factor Targets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| ZNF354A_TARGET_GENES | 6.69e-03 | 17.94 | 2.02 | 6.39e-01 | 1.00e+00 | 2RAI14, DLL3 |
31 |
| TAL1ALPHAE47_01 | 3.23e-03 | 5.49 | 1.69 | 6.39e-01 | 1.00e+00 | 5ASB4, HES6, NEUROD2, GFRA1, CLVS1 |
258 |
| SIX1_TARGET_GENES | 3.14e-03 | 4.61 | 1.59 | 6.39e-01 | 1.00e+00 | 6SIX1, HES6, ANO3, NKAIN4, AFAP1, NAV2 |
376 |
| AAANWWTGC_UNKNOWN | 7.07e-03 | 5.69 | 1.47 | 6.39e-01 | 1.00e+00 | 4LSAMP, SATB2, NNAT, FZD7 |
195 |
| CDP_02 | 9.53e-03 | 7.39 | 1.45 | 6.39e-01 | 1.00e+00 | 3SIX1, DACH1, LSAMP |
111 |
| HFH4_01 | 8.38e-03 | 5.41 | 1.40 | 6.39e-01 | 1.00e+00 | 4CYP26A1, NEUROD2, GFRA1, PCDH17 |
205 |
| TGACAGNY_MEIS1_01 | 3.80e-03 | 3.26 | 1.39 | 6.39e-01 | 1.00e+00 | 9CHRNA1, EYA2, HES6, NEUROD2, LSAMP, GIPR, PCDH17, NAV2, NNAT |
850 |
| RYTGCNWTGGNR_UNKNOWN | 1.12e-02 | 6.94 | 1.36 | 6.39e-01 | 1.00e+00 | 3SIX1, GRIN2D, NNAT |
118 |
| CAGCTG_AP4_Q5 | 9.36e-03 | 2.60 | 1.24 | 6.39e-01 | 1.00e+00 | 12LDB3, CHRNA1, EYA2, RAPSN, HES6, CERKL, NEUROD2, ASS1, GRIN2D, CLVS1, DLL3, NNAT |
1530 |
| ZF5_B | 1.48e-02 | 4.55 | 1.18 | 6.39e-01 | 1.00e+00 | 4CYP26A1, GFRA1, DDX1, NAV2 |
243 |
| PTF1BETA_Q6 | 1.54e-02 | 4.49 | 1.17 | 6.39e-01 | 1.00e+00 | 4ST18, CLVS1, PCDH17, ROBO1 |
246 |
| MYOD_Q6 | 1.60e-02 | 4.44 | 1.15 | 6.39e-01 | 1.00e+00 | 4CYP26A1, HES6, NKAIN4, GFRA1 |
249 |
| TAL1BETAE47_01 | 1.73e-02 | 4.33 | 1.12 | 6.39e-01 | 1.00e+00 | 4ASB4, NEUROD2, GFRA1, CLVS1 |
255 |
| AP4_Q6_01 | 1.77e-02 | 4.30 | 1.11 | 6.39e-01 | 1.00e+00 | 4RAPSN, HES6, GRIN2D, CLVS1 |
257 |
| ZA_UNIPROT_Q9UM89_UNREVIEWED_TARGET_GENES | 2.14e-02 | 9.46 | 1.09 | 6.39e-01 | 1.00e+00 | 2GKAP1, RAI14 |
57 |
| AP4_01 | 1.96e-02 | 4.16 | 1.08 | 6.39e-01 | 1.00e+00 | 4RAPSN, HES6, NEUROD2, GRIN2D |
265 |
| DBP_Q6 | 1.96e-02 | 4.16 | 1.08 | 6.39e-01 | 1.00e+00 | 4NEUROD2, GFRA1, GIPR, PCDH17 |
265 |
| E12_Q6 | 1.96e-02 | 4.16 | 1.08 | 6.39e-01 | 1.00e+00 | 4CYP26A1, HES6, GFRA1, DLL3 |
265 |
| TAL1BETAITF2_01 | 1.98e-02 | 4.15 | 1.08 | 6.39e-01 | 1.00e+00 | 4ASB4, NEUROD2, GFRA1, CLVS1 |
266 |
| OCT1_Q6 | 2.13e-02 | 4.06 | 1.05 | 6.39e-01 | 1.00e+00 | 4SIX1, NEUROD2, GNA14, SATB2 |
272 |
Dowload full table
GO Biological Processes:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| GOBP_ORGAN_INDUCTION | 3.10e-03 | 27.37 | 3.01 | 1.00e+00 | 1.00e+00 | 2SIX1, ROBO1 |
21 |
| GOBP_PROTEASOMAL_UBIQUITIN_INDEPENDENT_PROTEIN_CATABOLIC_PROCESS | 3.40e-03 | 26.01 | 2.87 | 1.00e+00 | 1.00e+00 | 2PSMB5, ENC1 |
22 |
| GOBP_RESPONSE_TO_NUTRIENT | 4.58e-04 | 8.68 | 2.65 | 1.00e+00 | 1.00e+00 | 5CYP26A1, ASS1, IGFBP2, GIPR, LDHA |
165 |
| GOBP_NEUROMUSCULAR_SYNAPTIC_TRANSMISSION | 4.05e-03 | 23.65 | 2.62 | 1.00e+00 | 1.00e+00 | 2CHRNA1, RAPSN |
24 |
| GOBP_REGULATION_OF_ANIMAL_ORGAN_FORMATION | 4.74e-03 | 21.66 | 2.42 | 1.00e+00 | 1.00e+00 | 2SIX1, ROBO1 |
26 |
| GOBP_SYNAPTIC_TRANSMISSION_CHOLINERGIC | 5.48e-03 | 20.00 | 2.24 | 1.00e+00 | 1.00e+00 | 2CHRNA1, RAPSN |
28 |
| GOBP_CELL_FATE_COMMITMENT_INVOLVED_IN_FORMATION_OF_PRIMARY_GERM_LAYER | 5.48e-03 | 20.00 | 2.24 | 1.00e+00 | 1.00e+00 | 2EYA2, FZD7 |
28 |
| GOBP_HEART_FORMATION | 5.87e-03 | 19.27 | 2.16 | 1.00e+00 | 1.00e+00 | 2SIX1, ROBO1 |
29 |
| GOBP_POSITIVE_REGULATION_OF_ANIMAL_ORGAN_MORPHOGENESIS | 6.27e-03 | 18.58 | 2.09 | 1.00e+00 | 1.00e+00 | 2SIX1, ROBO1 |
30 |
| GOBP_DEVELOPMENTAL_INDUCTION | 7.12e-03 | 17.34 | 1.95 | 1.00e+00 | 1.00e+00 | 2SIX1, ROBO1 |
32 |
| GOBP_SPECIFICATION_OF_ANIMAL_ORGAN_IDENTITY | 8.00e-03 | 16.26 | 1.84 | 1.00e+00 | 1.00e+00 | 2SIX1, ROBO1 |
34 |
| GOBP_CELL_FATE_SPECIFICATION | 5.05e-03 | 9.38 | 1.83 | 1.00e+00 | 1.00e+00 | 3SIX1, EYA2, FZD7 |
88 |
| GOBP_NEGATIVE_REGULATION_OF_PROTEASOMAL_UBIQUITIN_DEPENDENT_PROTEIN_CATABOLIC_PROCESS | 8.94e-03 | 15.30 | 1.73 | 1.00e+00 | 1.00e+00 | 2FHIT, SVIP |
36 |
| GOBP_REGULATION_OF_CAMP_MEDIATED_SIGNALING | 9.43e-03 | 14.87 | 1.69 | 1.00e+00 | 1.00e+00 | 2PEX5L, GIPR |
37 |
| GOBP_REGULATION_OF_ANIMAL_ORGAN_MORPHOGENESIS | 4.96e-03 | 6.32 | 1.63 | 1.00e+00 | 1.00e+00 | 4SIX1, ROBO1, FZD7, PSMB5 |
176 |
| GOBP_RESPONSE_TO_RETINOIC_ACID | 7.78e-03 | 7.98 | 1.56 | 1.00e+00 | 1.00e+00 | 3CYP26A1, IGFBP2, FZD7 |
103 |
| GOBP_NEUROMUSCULAR_JUNCTION_DEVELOPMENT | 1.49e-02 | 11.57 | 1.32 | 1.00e+00 | 1.00e+00 | 2SIX1, CHRNA1 |
47 |
| GOBP_RESPONSE_TO_ESTRADIOL | 1.31e-02 | 6.54 | 1.28 | 1.00e+00 | 1.00e+00 | 3SSTR2, ASS1, IGFBP2 |
125 |
| GOBP_ARGININE_BIOSYNTHETIC_PROCESS | 1.98e-02 | 63.54 | 1.27 | 1.00e+00 | 1.00e+00 | 1ASS1 |
5 |
| GOBP_NEGATIVE_REGULATION_OF_CELL_FATE_SPECIFICATION | 1.98e-02 | 63.54 | 1.27 | 1.00e+00 | 1.00e+00 | 1FZD7 |
5 |
Dowload full table
Immunological Gene Sets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| GSE17721_0.5H_VS_8H_GARDIQUIMOD_BMDC_DN | 1.06e-03 | 7.16 | 2.19 | 1.00e+00 | 1.00e+00 | 5DACH1, PAPSS1, DLK1, SATB2, FZD7 |
199 |
| GSE15659_NONSUPPRESSIVE_TCELL_VS_ACTIVATED_TREG_UP | 6.95e-03 | 5.72 | 1.48 | 1.00e+00 | 1.00e+00 | 4ASB4, AFAP1, GFRA1, DLL3 |
194 |
| GSE29618_PRE_VS_DAY7_POST_LAIV_FLU_VACCINE_PDC_UP | 7.20e-03 | 5.66 | 1.47 | 1.00e+00 | 1.00e+00 | 4RAI14, AFAP1, GIPR, ROBO1 |
196 |
| GSE21546_UNSTIM_VS_ANTI_CD3_STIM_SAP1A_KO_DP_THYMOCYTES_UP | 7.45e-03 | 5.60 | 1.45 | 1.00e+00 | 1.00e+00 | 4CYP26A1, PAPSS1, RAI14, GFRA1 |
198 |
| GSE21927_C26GM_VS_4T1_TUMOR_MONOCYTE_BALBC_UP | 7.45e-03 | 5.60 | 1.45 | 1.00e+00 | 1.00e+00 | 4PPP1R14A, HES6, PAPSS1, GRIN2D |
198 |
| GSE29949_MICROGLIA_BRAIN_VS_CD8_NEG_DC_SPLEEN_DN | 7.45e-03 | 5.60 | 1.45 | 1.00e+00 | 1.00e+00 | 4NEUROD2, DLK1, SMAD9, KCNJ6 |
198 |
| GSE17721_CTRL_VS_LPS_1H_BMDC_DN | 7.58e-03 | 5.57 | 1.44 | 1.00e+00 | 1.00e+00 | 4SIX1, CYP26A1, RAI14, CLVS1 |
199 |
| GSE17721_CTRL_VS_CPG_24H_BMDC_UP | 7.71e-03 | 5.55 | 1.44 | 1.00e+00 | 1.00e+00 | 4EYA2, HES6, GNA14, DLL3 |
200 |
| GSE17721_CPG_VS_GARDIQUIMOD_12H_BMDC_DN | 7.71e-03 | 5.55 | 1.44 | 1.00e+00 | 1.00e+00 | 4SIX1, DACH1, RAPSN, SATB2 |
200 |
| GSE9878_CTRL_VS_EBF_TRANSDUCED_PAX5_KO_PRO_BCELL_UP | 7.71e-03 | 5.55 | 1.44 | 1.00e+00 | 1.00e+00 | 4EYA2, GKAP1, SVIP, RBFOX3 |
200 |
| GSE21670_UNTREATED_VS_TGFB_TREATED_CD4_TCELL_UP | 7.71e-03 | 5.55 | 1.44 | 1.00e+00 | 1.00e+00 | 4ASS1, DLK1, SMAD9, KCNJ6 |
200 |
| GSE29949_MICROGLIA_BRAIN_VS_CD8_POS_DC_SPLEEN_DN | 7.71e-03 | 5.55 | 1.44 | 1.00e+00 | 1.00e+00 | 4LDB3, NEUROD2, SATB2, FZD7 |
200 |
| GSE11961_GERMINAL_CENTER_BCELL_DAY7_VS_MEMORY_BCELL_DAY40_DN | 7.71e-03 | 5.55 | 1.44 | 1.00e+00 | 1.00e+00 | 4SSTR2, PPP1R14A, THSD7B, GFRA1 |
200 |
| GSE13547_WT_VS_ZFX_KO_BCELL_ANTI_IGM_STIM_12H_UP | 2.00e-02 | 5.54 | 1.09 | 1.00e+00 | 1.00e+00 | 3DACH1, PCDH17, KCNJ6 |
147 |
| GSE9946_IMMATURE_VS_PROSTAGLANDINE2_TREATED_MATURE_DC_UP | 2.08e-02 | 5.46 | 1.08 | 1.00e+00 | 1.00e+00 | 3SSTR2, LSAMP, ANO3 |
149 |
| GSE13522_CTRL_VS_T_CRUZI_Y_STRAIN_INF_SKIN_IFNG_KO_UP | 2.38e-02 | 5.18 | 1.02 | 1.00e+00 | 1.00e+00 | 3RAPSN, ANO3, FZD7 |
157 |
| GSE32255_WT_UNSTIM_VS_JMJD2D_KNOCKDOWN_4H_LPS_STIM_DC_DN | 3.04e-02 | 4.69 | 0.93 | 1.00e+00 | 1.00e+00 | 3RAPSN, NAV2, STARD4 |
173 |
| GSE36095_WT_VS_HDAC9_KO_TREG_UP | 3.40e-02 | 4.48 | 0.88 | 1.00e+00 | 1.00e+00 | 3EYA2, FAM78B, LDHA |
181 |
| GSE16385_UNTREATED_VS_12H_ROSIGLITAZONE_TREATED_MACROPHAGE_UP | 3.74e-02 | 4.31 | 0.85 | 1.00e+00 | 1.00e+00 | 3RAI14, NKAIN4, ENC1 |
188 |
| GSE45365_BCELL_VS_CD8_TCELL_DN | 3.74e-02 | 4.31 | 0.85 | 1.00e+00 | 1.00e+00 | 3PPP1R14A, IGFBP2, GNA14 |
188 |
Top Ranked Transcription Factors for this Gene Expression Program:
| Gene Symbol | TF Rank | DNA Binding Domain | Motif Status | IUPAC PWM | GTEx | DepMap | Decartes |
|---|---|---|---|---|---|---|---|
| SIX1 | 2 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| DACH1 | 6 | Yes | Likely to be sequence specific TF | Monomer or homomultimer | No motif | Has a putative AT-hook | Binds with consensus AAWANAAAWAAWT and AATACAATTAAAT as strongest target sequences based on EMSA and SELEX (PMID: 20351289). Protein contains a winged helix structural domain (PMID: 12057194) |
| HES6 | 12 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| NEUROD2 | 16 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| ST18 | 17 | Yes | Inferred motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| SMAD9 | 21 | Yes | Known motif | Obligate heteromer | In vivo/Misc source | Only known motifs are from Transfac or HocoMoco - origin is uncertain | None |
| SATB2 | 36 | Yes | Inferred motif | Low specificity DNA-binding protein | In vivo/Misc source | None | SATBs were analyzed by Gwenael Badis and Mike Berger a decade ago; by PBMs. It did not yield a motif; instead; the signal was very closely proportional to nucleotide content, as the name suggests (Special AT Binding). |
| ZBTB18 | 59 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| CYP1B1 | 63 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | None |
| TCF15 | 66 | Yes | Inferred motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| NRIP1 | 67 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | Transcriptional cofactor |
| HHEX | 91 | Yes | Inferred motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| RCOR2 | 95 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | Contains 2 SANT domains, and no other putative DNA-binding domains |
| PAX6 | 103 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| NPM1 | 109 | No | Unlikely to be sequence specific TF | Low specificity DNA-binding protein | No motif | None | No evidence for sequence-specific DNA-binding (PMID: 2223875) |
| TSN | 111 | No | Unlikely to be sequence specific TF | Low specificity DNA-binding protein | No motif | None | Based on (PMID: 20450889) and (PMID: 12484770), this protein binds repetitive DNA sequences as multimeric complexes that may contain partner protein TSNAX |
| H1FX | 117 | No | Unlikely to be sequence specific TF | Low specificity DNA-binding protein | No motif | None | Histone component |
| DACH2 | 119 | Yes | Likely to be sequence specific TF | Monomer or homomultimer | No motif | Has a putative AT-hook | Related to DACH1, which can bind DNA based on EMSA and SELEX (PMID: 20351289). Yet, the fly ortholog has been extensively studied and cannot bind DNA. (PMID: 8431945; PMID: 7821215) |
| CNBP | 131 | No | ssDNA/RNA binding | Not a DNA binding protein | No motif | None | CNBP has a specificity for single-stranded DNA (PMID: 2562787) |
| SETBP1 | 133 | Yes | Likely to be sequence specific TF | Monomer or homomultimer | No motif | None | Orthologous protein from mouse (Setbp1) bind DNA sequence-specifically by PBM |
QQ Plot showing correlations with other GEPs in this dataset, calculated by Spearman correlation:
Interactive QQ-plot of gene loadings:
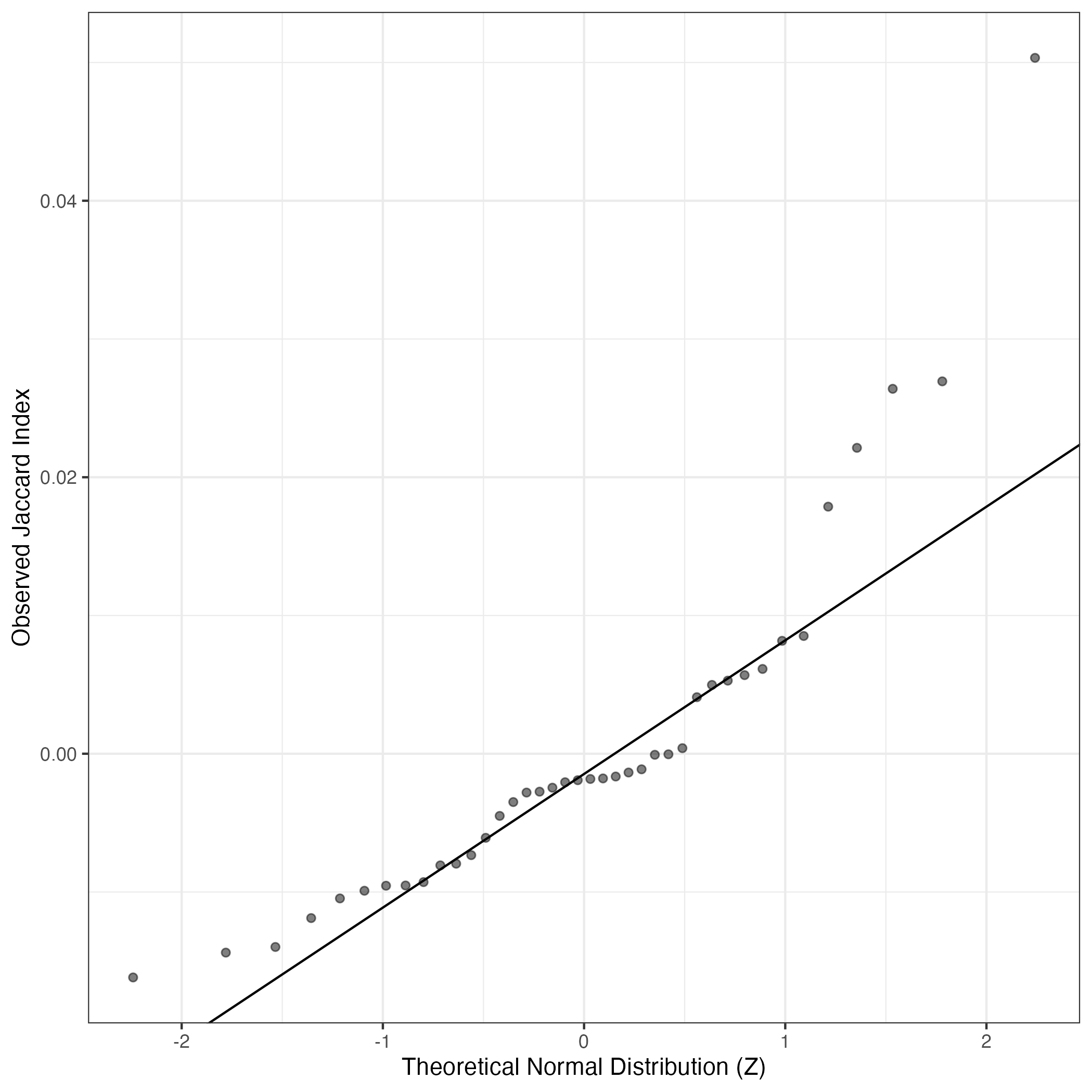
A similar QQ-plot as above, but only for instances where the H value is e.g. > 25, i.e. we are confident that the expression program is active above noise. Agreemenet between these binary vectors is tested using the Jaccard Index, with the P-values calculated by an exact test:
Interactive QQ-plot:
Singler cell type annotations for the top 50 cells on this program.
| Cell ID | Singler label | Singler Delta | Activity Score | Top Singler Raw Scores |
|---|---|---|---|---|
| T162_AAGATAGCAACTTGCA-1 | Neurons:adrenal_medulla_cell_line | 0.17 | 663.40 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.35, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_1lox-17Puro-10: 0.33, iPS_cells:PDB_1lox-17Puro-5: 0.33, iPS_cells:PDB_2lox-17: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.33, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.32 |
| T162_CTGTGAACACGCTTAA-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 513.94 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.36, Neuroepithelial_cell:ESC-derived: 0.34, iPS_cells:PDB_2lox-22: 0.32, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:PDB_2lox-5: 0.32, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32 |
| T162_CTGTACCAGCCATTCA-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 401.60 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.38, Neuroepithelial_cell:ESC-derived: 0.34, Astrocyte:Embryonic_stem_cell-derived: 0.32, iPS_cells:PDB_2lox-22: 0.31, iPS_cells:PDB_1lox-21Puro-20: 0.31, iPS_cells:PDB_2lox-5: 0.31, Neurons:ES_cell-derived_neural_precursor: 0.31, iPS_cells:PDB_1lox-21Puro-26: 0.31, iPS_cells:PDB_1lox-17Puro-10: 0.31, iPS_cells:PDB_2lox-17: 0.31 |
| T162_ATGGAGGCACAACATC-1 | Neurons:adrenal_medulla_cell_line | 0.17 | 395.53 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.4, Neuroepithelial_cell:ESC-derived: 0.39, iPS_cells:PDB_1lox-21Puro-20: 0.36, iPS_cells:PDB_2lox-22: 0.36, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.36, Astrocyte:Embryonic_stem_cell-derived: 0.36, iPS_cells:PDB_1lox-17Puro-10: 0.36, iPS_cells:PDB_1lox-21Puro-26: 0.36, iPS_cells:PDB_2lox-21: 0.35, iPS_cells:PDB_2lox-17: 0.35 |
| T162_AGCATCATCGAAACAA-1 | Neurons:adrenal_medulla_cell_line | 0.20 | 394.42 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.4, Neuroepithelial_cell:ESC-derived: 0.36, Astrocyte:Embryonic_stem_cell-derived: 0.34, iPS_cells:PDB_1lox-17Puro-10: 0.33, iPS_cells:PDB_2lox-22: 0.33, Embryonic_stem_cells: 0.33, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_2lox-5: 0.32 |
| T162_GATCCCTAGCACCTGC-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 343.91 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.36, Neuroepithelial_cell:ESC-derived: 0.35, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.32, iPS_cells:PDB_2lox-22: 0.32, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32, iPS_cells:PDB_1lox-17Puro-5: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.32 |
| T162_AGTCAACAGTCTACCA-1 | Neurons:adrenal_medulla_cell_line | 0.16 | 334.04 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.36, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_2lox-21: 0.33, iPS_cells:PDB_1lox-17Puro-10: 0.33, iPS_cells:PDB_1lox-17Puro-5: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.33, iPS_cells:PDB_2lox-17: 0.33, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.33 |
| T162_CGAATTGGTGGAACCA-1 | Neurons:adrenal_medulla_cell_line | 0.21 | 323.54 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.41, Neuroepithelial_cell:ESC-derived: 0.36, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.33, iPS_cells:PDB_1lox-17Puro-10: 0.33, iPS_cells:PDB_2lox-21: 0.33, iPS_cells:PDB_2lox-17: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.33, iPS_cells:PDB_1lox-17Puro-5: 0.33 |
| T162_TTCCTAATCTCTGCTG-1 | Neurons:adrenal_medulla_cell_line | 0.17 | 322.06 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.4, Neuroepithelial_cell:ESC-derived: 0.38, iPS_cells:PDB_1lox-21Puro-20: 0.37, iPS_cells:PDB_1lox-21Puro-26: 0.36, iPS_cells:PDB_1lox-17Puro-10: 0.36, iPS_cells:PDB_2lox-22: 0.36, iPS_cells:PDB_1lox-17Puro-5: 0.36, iPS_cells:PDB_2lox-21: 0.36, iPS_cells:PDB_2lox-17: 0.36, Embryonic_stem_cells: 0.36 |
| T162_CCAATGACAGCTTCCT-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 320.07 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.35, Neuroepithelial_cell:ESC-derived: 0.32, iPS_cells:PDB_2lox-22: 0.3, iPS_cells:PDB_1lox-21Puro-20: 0.3, iPS_cells:PDB_2lox-21: 0.3, iPS_cells:PDB_1lox-21Puro-26: 0.3, iPS_cells:PDB_2lox-17: 0.3, iPS_cells:PDB_1lox-17Puro-10: 0.3, iPS_cells:PDB_2lox-5: 0.3, iPS_cells:PDB_1lox-17Puro-5: 0.3 |
| T162_GATTCGAAGTCGGCAA-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 316.51 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.41, Neuroepithelial_cell:ESC-derived: 0.38, iPS_cells:PDB_1lox-21Puro-20: 0.36, iPS_cells:PDB_1lox-17Puro-5: 0.36, iPS_cells:PDB_1lox-17Puro-10: 0.36, iPS_cells:PDB_2lox-22: 0.36, iPS_cells:PDB_1lox-21Puro-26: 0.35, iPS_cells:PDB_2lox-21: 0.35, iPS_cells:PDB_2lox-17: 0.35, Astrocyte:Embryonic_stem_cell-derived: 0.35 |
| T162_ACGTACAAGTGCCCGT-1 | Neurons:adrenal_medulla_cell_line | 0.18 | 313.69 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.38, Neuroepithelial_cell:ESC-derived: 0.33, iPS_cells:PDB_2lox-22: 0.31, Astrocyte:Embryonic_stem_cell-derived: 0.31, iPS_cells:PDB_2lox-21: 0.3, iPS_cells:PDB_2lox-17: 0.3, iPS_cells:PDB_1lox-21Puro-20: 0.3, iPS_cells:PDB_1lox-17Puro-10: 0.3, iPS_cells:PDB_1lox-21Puro-26: 0.3, iPS_cells:PDB_1lox-17Puro-5: 0.3 |
| T162_TGTCAGAAGGTTACAA-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 299.30 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.32, Neuroepithelial_cell:ESC-derived: 0.28, Astrocyte:Embryonic_stem_cell-derived: 0.26, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.26, iPS_cells:PDB_1lox-21Puro-20: 0.26, iPS_cells:PDB_1lox-17Puro-10: 0.26, iPS_cells:PDB_1lox-21Puro-26: 0.26, iPS_cells:PDB_1lox-17Puro-5: 0.26, iPS_cells:PDB_2lox-22: 0.25, iPS_cells:PDB_2lox-17: 0.25 |
| T162_AACCAACTCAATCAGC-1 | Neurons:adrenal_medulla_cell_line | 0.20 | 293.54 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.34, iPS_cells:PDB_2lox-22: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.32, iPS_cells:PDB_1lox-17Puro-5: 0.31, iPS_cells:PDB_2lox-17: 0.31, iPS_cells:PDB_2lox-21: 0.31, Embryonic_stem_cells: 0.31 |
| T162_GATTTCTTCTGGCCGA-1 | Neurons:adrenal_medulla_cell_line | 0.16 | 292.77 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.36, Neuroepithelial_cell:ESC-derived: 0.34, iPS_cells:PDB_2lox-22: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.31, iPS_cells:PDB_2lox-21: 0.31, iPS_cells:PDB_1lox-17Puro-10: 0.31, Astrocyte:Embryonic_stem_cell-derived: 0.31, iPS_cells:PDB_1lox-21Puro-26: 0.31 |
| T162_AACAAAGTCGTGCGAC-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 283.57 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.39, Neuroepithelial_cell:ESC-derived: 0.34, iPS_cells:PDB_2lox-22: 0.32, Astrocyte:Embryonic_stem_cell-derived: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_2lox-21: 0.31, iPS_cells:PDB_1lox-21Puro-20: 0.31, iPS_cells:PDB_1lox-17Puro-10: 0.31, iPS_cells:PDB_1lox-21Puro-26: 0.31, iPS_cells:PDB_2lox-5: 0.31 |
| T162_AGCATCATCCATTTAC-1 | Neurons:adrenal_medulla_cell_line | 0.16 | 276.84 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.34, Astrocyte:Embryonic_stem_cell-derived: 0.33, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.32, iPS_cells:PDB_2lox-22: 0.32, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:iPS:minicircle-derived: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32 |
| T162_AGTACCAAGAAACCCG-1 | Neurons:adrenal_medulla_cell_line | 0.17 | 275.08 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.36, Neuroepithelial_cell:ESC-derived: 0.35, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_1lox-17Puro-10: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.33, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.33, iPS_cells:PDB_1lox-17Puro-5: 0.33, iPS_cells:PDB_2lox-21: 0.33, iPS_cells:PDB_2lox-17: 0.33 |
| T162_GGCACGTTCATGCCCT-1 | Neurons:adrenal_medulla_cell_line | 0.11 | 273.21 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.28, Neuroepithelial_cell:ESC-derived: 0.26, iPS_cells:PDB_1lox-17Puro-10: 0.26, iPS_cells:PDB_1lox-17Puro-5: 0.26, iPS_cells:PDB_2lox-22: 0.26, iPS_cells:PDB_2lox-17: 0.26, iPS_cells:PDB_1lox-21Puro-26: 0.26, iPS_cells:PDB_2lox-21: 0.26, iPS_cells:PDB_1lox-21Puro-20: 0.26, iPS_cells:PDB_2lox-5: 0.26 |
| T162_CGGTCAGTCTGATTCT-1 | Neurons:adrenal_medulla_cell_line | 0.18 | 267.37 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.35, Neuroepithelial_cell:ESC-derived: 0.31, Astrocyte:Embryonic_stem_cell-derived: 0.29, iPS_cells:PDB_1lox-21Puro-20: 0.28, iPS_cells:PDB_2lox-22: 0.28, iPS_cells:PDB_1lox-17Puro-10: 0.28, iPS_cells:PDB_1lox-17Puro-5: 0.28, iPS_cells:PDB_1lox-21Puro-26: 0.28, Neurons:ES_cell-derived_neural_precursor: 0.27, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.27 |
| T162_TTCATTGTCTCCACTG-1 | Neurons:adrenal_medulla_cell_line | 0.17 | 260.29 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.35, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:PDB_1lox-17Puro-5: 0.32, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.32 |
| T162_AGTCACAAGACATAAC-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 245.18 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.4, Neuroepithelial_cell:ESC-derived: 0.36, Astrocyte:Embryonic_stem_cell-derived: 0.34, iPS_cells:PDB_1lox-21Puro-20: 0.34, iPS_cells:PDB_1lox-17Puro-10: 0.34, iPS_cells:PDB_2lox-22: 0.34, iPS_cells:PDB_1lox-21Puro-26: 0.34, iPS_cells:PDB_1lox-17Puro-5: 0.34, iPS_cells:PDB_2lox-21: 0.33, iPS_cells:PDB_2lox-17: 0.33 |
| T162_CCTCCTCAGCGAAACC-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 234.60 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.29, Neuroepithelial_cell:ESC-derived: 0.26, Astrocyte:Embryonic_stem_cell-derived: 0.24, Embryonic_stem_cells: 0.23, Neurons:ES_cell-derived_neural_precursor: 0.23, iPS_cells:PDB_1lox-17Puro-10: 0.22, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.22, iPS_cells:PDB_2lox-22: 0.22, iPS_cells:PDB_1lox-17Puro-5: 0.22, iPS_cells:PDB_1lox-21Puro-20: 0.22 |
| T162_GGGTGAACAACTCCCT-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 234.52 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.4, Neuroepithelial_cell:ESC-derived: 0.37, iPS_cells:PDB_1lox-21Puro-20: 0.35, iPS_cells:PDB_1lox-21Puro-26: 0.34, iPS_cells:PDB_2lox-22: 0.34, iPS_cells:PDB_1lox-17Puro-10: 0.34, iPS_cells:PDB_1lox-17Puro-5: 0.34, Astrocyte:Embryonic_stem_cell-derived: 0.34, iPS_cells:PDB_2lox-17: 0.34, iPS_cells:PDB_2lox-21: 0.34 |
| T162_GATGGAGGTTAATGAG-1 | Neurons:adrenal_medulla_cell_line | 0.16 | 234.15 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.38, Neuroepithelial_cell:ESC-derived: 0.35, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_2lox-22: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.33, Embryonic_stem_cells: 0.33, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.33, iPS_cells:PDB_1lox-17Puro-10: 0.33, iPS_cells:PDB_2lox-21: 0.33 |
| T162_GCAGGCTTCCTAAACG-1 | Neurons:adrenal_medulla_cell_line | 0.20 | 233.14 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.38, Neuroepithelial_cell:ESC-derived: 0.34, Astrocyte:Embryonic_stem_cell-derived: 0.33, iPS_cells:PDB_1lox-21Puro-20: 0.31, iPS_cells:PDB_1lox-17Puro-10: 0.31, iPS_cells:PDB_2lox-22: 0.31, iPS_cells:PDB_1lox-17Puro-5: 0.31, iPS_cells:PDB_1lox-21Puro-26: 0.31, iPS_cells:PDB_2lox-21: 0.31, iPS_cells:PDB_2lox-17: 0.31 |
| T162_GGTCTGGTCACCATCC-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 230.91 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.34, Astrocyte:Embryonic_stem_cell-derived: 0.33, iPS_cells:PDB_2lox-22: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.31, iPS_cells:PDB_1lox-17Puro-5: 0.31 |
| T162_TGGTTAGCAATCGTCA-1 | Monocyte:MCSF | 0.03 | 229.49 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.34, Neuroepithelial_cell:ESC-derived: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.33, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.31, Embryonic_stem_cells: 0.3, iPS_cells:PDB_2lox-22: 0.3, Neurons:Schwann_cell: 0.3, iPS_cells:PDB_2lox-5: 0.3, iPS_cells:fibroblast-derived:Retroviral_transf: 0.3, iPS_cells:PDB_2lox-17: 0.3 |
| T162_TCGGGACCAGCCTTCT-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 228.84 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.35, Neuroepithelial_cell:ESC-derived: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.31, iPS_cells:PDB_1lox-21Puro-20: 0.31, iPS_cells:PDB_1lox-17Puro-10: 0.31, iPS_cells:PDB_2lox-22: 0.31, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.31, iPS_cells:PDB_2lox-21: 0.31, iPS_cells:PDB_2lox-17: 0.31, iPS_cells:PDB_1lox-21Puro-26: 0.31 |
| T162_TTGTTTGTCGCAACAT-1 | Neurons:adrenal_medulla_cell_line | 0.20 | 226.43 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.29, Embryonic_stem_cells: 0.29, Neurons:ES_cell-derived_neural_precursor: 0.29, iPS_cells:PDB_1lox-17Puro-10: 0.29, iPS_cells:PDB_2lox-22: 0.29, iPS_cells:PDB_1lox-21Puro-26: 0.29, iPS_cells:PDB_1lox-17Puro-5: 0.29 |
| T162_ACTTCCGTCCGTAGTA-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 216.55 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.36, Neuroepithelial_cell:ESC-derived: 0.35, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.32, Embryonic_stem_cells: 0.32, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32 |
| T162_TGTTCATGTCTTACAG-1 | Neurons:adrenal_medulla_cell_line | 0.17 | 214.86 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.38, Neuroepithelial_cell:ESC-derived: 0.35, iPS_cells:PDB_2lox-22: 0.34, iPS_cells:PDB_2lox-17: 0.33, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_2lox-21: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.33, Embryonic_stem_cells: 0.33, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.33 |
| T162_ACCTGAAGTGACACGA-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 212.22 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.31, Neuroepithelial_cell:ESC-derived: 0.29, iPS_cells:PDB_2lox-22: 0.27, Astrocyte:Embryonic_stem_cell-derived: 0.27, iPS_cells:PDB_2lox-21: 0.27, iPS_cells:PDB_1lox-21Puro-20: 0.27, iPS_cells:PDB_2lox-17: 0.27, iPS_cells:PDB_2lox-5: 0.27, iPS_cells:PDB_1lox-17Puro-10: 0.26, iPS_cells:PDB_1lox-21Puro-26: 0.26 |
| T162_CCGTAGGTCGCAACAT-1 | Neurons:adrenal_medulla_cell_line | 0.18 | 210.15 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.35, Neuroepithelial_cell:ESC-derived: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.31, iPS_cells:PDB_2lox-22: 0.29, Neurons:ES_cell-derived_neural_precursor: 0.29, iPS_cells:PDB_1lox-21Puro-20: 0.29, iPS_cells:PDB_1lox-17Puro-10: 0.29, iPS_cells:PDB_1lox-21Puro-26: 0.29, iPS_cells:PDB_2lox-21: 0.29, iPS_cells:PDB_2lox-17: 0.29 |
| T162_TGCAGTAAGCTTCTAG-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 209.12 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.3, Neuroepithelial_cell:ESC-derived: 0.27, Astrocyte:Embryonic_stem_cell-derived: 0.26, iPS_cells:PDB_2lox-22: 0.24, iPS_cells:PDB_2lox-17: 0.24, iPS_cells:PDB_1lox-17Puro-5: 0.24, iPS_cells:PDB_2lox-21: 0.24, iPS_cells:PDB_1lox-17Puro-10: 0.24, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.24, iPS_cells:PDB_2lox-5: 0.24 |
| T162_AGCTACAAGGTGGCTA-1 | Neurons:adrenal_medulla_cell_line | 0.18 | 203.33 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.36, Neuroepithelial_cell:ESC-derived: 0.31, Astrocyte:Embryonic_stem_cell-derived: 0.29, iPS_cells:PDB_2lox-22: 0.28, Neurons:ES_cell-derived_neural_precursor: 0.28, iPS_cells:PDB_1lox-21Puro-20: 0.28, iPS_cells:PDB_2lox-21: 0.27, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.27, iPS_cells:PDB_1lox-21Puro-26: 0.27, iPS_cells:PDB_2lox-17: 0.27 |
| T162_TTCCAATCAAGTATAG-1 | Neurons:adrenal_medulla_cell_line | 0.11 | 198.83 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.26, Neuroepithelial_cell:ESC-derived: 0.24, iPS_cells:adipose_stem_cell-derived:minicircle-derived: 0.23, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.23, Astrocyte:Embryonic_stem_cell-derived: 0.23, iPS_cells:adipose_stem_cell-derived:lentiviral: 0.23, iPS_cells:PDB_2lox-22: 0.22, iPS_cells:PDB_2lox-21: 0.22, Embryonic_stem_cells: 0.22, iPS_cells:PDB_2lox-5: 0.22 |
| T162_GGGTAGAGTCTGGTTA-1 | Neurons:adrenal_medulla_cell_line | 0.18 | 194.36 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.4, Neuroepithelial_cell:ESC-derived: 0.35, Astrocyte:Embryonic_stem_cell-derived: 0.33, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_2lox-21: 0.32, iPS_cells:PDB_2lox-17: 0.32, iPS_cells:adipose_stem_cell-derived:minicircle-derived: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.32 |
| T162_AGTGACTAGGTGCTAG-1 | Neurons:adrenal_medulla_cell_line | 0.16 | 193.13 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.34, Neuroepithelial_cell:ESC-derived: 0.3, Astrocyte:Embryonic_stem_cell-derived: 0.29, iPS_cells:PDB_2lox-22: 0.27, iPS_cells:PDB_2lox-5: 0.27, Embryonic_stem_cells: 0.27, iPS_cells:PDB_1lox-17Puro-10: 0.27, iPS_cells:PDB_2lox-17: 0.27, iPS_cells:PDB_1lox-21Puro-20: 0.27, iPS_cells:PDB_2lox-21: 0.27 |
| T162_AAGTTCGGTATCCCTC-1 | Neurons:adrenal_medulla_cell_line | 0.09 | 191.94 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.3, Neuroepithelial_cell:ESC-derived: 0.27, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.26, Astrocyte:Embryonic_stem_cell-derived: 0.26, iPS_cells:PDB_1lox-17Puro-5: 0.26, iPS_cells:PDB_2lox-22: 0.26, iPS_cells:PDB_1lox-17Puro-10: 0.26, iPS_cells:PDB_1lox-21Puro-26: 0.26, iPS_cells:PDB_1lox-21Puro-20: 0.26, iPS_cells:PDB_2lox-17: 0.26 |
| T162_AATCACGAGGCTTTCA-1 | Neurons:adrenal_medulla_cell_line | 0.12 | 184.31 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.28, Neuroepithelial_cell:ESC-derived: 0.25, Astrocyte:Embryonic_stem_cell-derived: 0.25, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.24, Embryonic_stem_cells: 0.24, iPS_cells:PDB_2lox-22: 0.23, iPS_cells:PDB_1lox-21Puro-20: 0.23, iPS_cells:PDB_2lox-5: 0.23, iPS_cells:PDB_1lox-17Puro-10: 0.23, iPS_cells:PDB_2lox-17: 0.23 |
| T162_GACTTCCTCTGTGCGG-1 | Neurons:adrenal_medulla_cell_line | 0.18 | 184.26 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.39, Neuroepithelial_cell:ESC-derived: 0.36, Astrocyte:Embryonic_stem_cell-derived: 0.34, iPS_cells:PDB_1lox-21Puro-20: 0.33, iPS_cells:PDB_2lox-22: 0.33, iPS_cells:PDB_1lox-21Puro-26: 0.33, iPS_cells:PDB_2lox-17: 0.33, iPS_cells:PDB_2lox-5: 0.33, iPS_cells:PDB_2lox-21: 0.33, iPS_cells:PDB_1lox-17Puro-10: 0.33 |
| T162_CAACGATAGATGCGAC-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 182.45 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.37, Neuroepithelial_cell:ESC-derived: 0.33, iPS_cells:PDB_2lox-22: 0.31, iPS_cells:PDB_1lox-21Puro-20: 0.31, iPS_cells:PDB_1lox-17Puro-10: 0.31, Astrocyte:Embryonic_stem_cell-derived: 0.31, iPS_cells:PDB_1lox-21Puro-26: 0.31, iPS_cells:PDB_1lox-17Puro-5: 0.31, iPS_cells:PDB_2lox-17: 0.31, iPS_cells:PDB_2lox-21: 0.31 |
| T162_TCCTCTTGTTCGGTCG-1 | Neurons:adrenal_medulla_cell_line | 0.15 | 182.19 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.34, Neuroepithelial_cell:ESC-derived: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.29, iPS_cells:PDB_2lox-22: 0.29, Astrocyte:Embryonic_stem_cell-derived: 0.29, iPS_cells:PDB_1lox-17Puro-10: 0.29, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.29, iPS_cells:PDB_1lox-21Puro-26: 0.29, iPS_cells:PDB_1lox-17Puro-5: 0.29, Embryonic_stem_cells: 0.29 |
| T162_CATACAGCATCGAAGG-1 | Neurons:adrenal_medulla_cell_line | 0.21 | 174.89 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.43, Neuroepithelial_cell:ESC-derived: 0.38, iPS_cells:PDB_2lox-22: 0.36, iPS_cells:PDB_1lox-21Puro-20: 0.36, iPS_cells:PDB_2lox-21: 0.36, iPS_cells:PDB_2lox-17: 0.36, iPS_cells:PDB_1lox-21Puro-26: 0.36, Neurons:ES_cell-derived_neural_precursor: 0.35, Astrocyte:Embryonic_stem_cell-derived: 0.35, iPS_cells:PDB_1lox-17Puro-10: 0.35 |
| T162_GGGTGAACAGCAGTAG-1 | Neurons:adrenal_medulla_cell_line | 0.20 | 174.14 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.4, Neuroepithelial_cell:ESC-derived: 0.36, iPS_cells:PDB_1lox-21Puro-20: 0.35, iPS_cells:PDB_1lox-21Puro-26: 0.35, iPS_cells:PDB_2lox-22: 0.35, iPS_cells:PDB_1lox-17Puro-10: 0.35, iPS_cells:PDB_2lox-17: 0.35, iPS_cells:PDB_2lox-21: 0.35, Embryonic_stem_cells: 0.34, iPS_cells:PDB_1lox-17Puro-5: 0.34 |
| T162_GGATCTAAGCAACAGC-1 | Neurons:adrenal_medulla_cell_line | 0.19 | 172.20 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.38, Neuroepithelial_cell:ESC-derived: 0.34, Astrocyte:Embryonic_stem_cell-derived: 0.32, iPS_cells:PDB_1lox-21Puro-20: 0.32, iPS_cells:PDB_1lox-17Puro-10: 0.32, iPS_cells:PDB_1lox-21Puro-26: 0.32, iPS_cells:PDB_1lox-17Puro-5: 0.32, iPS_cells:PDB_2lox-22: 0.31, iPS_cells:PDB_2lox-17: 0.31, iPS_cells:PDB_2lox-21: 0.31 |
| T162_ATGAGTCAGCAGCCTC-1 | Neurons:adrenal_medulla_cell_line | 0.11 | 172.05 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.29, Neuroepithelial_cell:ESC-derived: 0.25, Astrocyte:Embryonic_stem_cell-derived: 0.25, iPS_cells:CRL2097_foreskin-derived:undiff.: 0.25, Embryonic_stem_cells: 0.24, iPS_cells:PDB_1lox-17Puro-5: 0.24, iPS_cells:PDB_2lox-22: 0.24, iPS_cells:PDB_1lox-21Puro-20: 0.24, iPS_cells:PDB_1lox-21Puro-26: 0.24, iPS_cells:PDB_2lox-17: 0.24 |
| T162_ACGTACATCGAACCTA-1 | Neurons:adrenal_medulla_cell_line | 0.17 | 171.55 | Raw ScoresNeurons:adrenal_medulla_cell_line: 0.36, Neuroepithelial_cell:ESC-derived: 0.33, Astrocyte:Embryonic_stem_cell-derived: 0.31, Neurons:ES_cell-derived_neural_precursor: 0.29, iPS_cells:PDB_2lox-22: 0.29, iPS_cells:PDB_1lox-21Puro-20: 0.29, iPS_cells:PDB_1lox-17Puro-10: 0.29, Embryonic_stem_cells: 0.28, iPS_cells:PDB_1lox-21Puro-26: 0.28, iPS_cells:PDB_2lox-21: 0.28 |
| T162_TCGGGACTCGCACGGT-1 | Macrophage:monocyte-derived:M-CSF | 0.07 | 171.39 | Raw ScoresDC:monocyte-derived:AEC-conditioned: 0.34, Macrophage:monocyte-derived: 0.34, Macrophage:monocyte-derived:IL-4/Dex/TGFb: 0.33, Macrophage:monocyte-derived:IL-4/Dex/cntrl: 0.33, Macrophage:monocyte-derived:M-CSF: 0.33, Monocyte:MCSF: 0.33, Macrophage:monocyte-derived:IL-4/cntrl: 0.33, Macrophage:monocyte-derived:IL-4/TGFb: 0.33, DC:monocyte-derived: 0.33, Macrophage:Alveolar: 0.33 |
Below shows the significant enrichments of this GEP for literature curated gene lists
This data was procured from existing single cell RNA-seq maps of neuroblastoma or related relevant data.
High ranks indicate this gene is a driver of this GEP.
These curated gene list are ranked by the P-value (on this GEP) of their constituent genes.
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
Adrenal Premordium (Hanemaaijer)
Marker genes obtained from Supplementary Table SD of Hanemaaijer et al (PMID 33500353). The authors generated single-cell RNA-seq data (sort-seq, 2,229 cells total) from mouse adrenal glads at E13.5, E14.5, E17.5, E18.5, P1 and P5. These were marker genes that matched with a similar dataset generated by Furlan et al (PMID 28684471). This particular set of markers are for the Adrenal Premordium subcluster, which is part of the Cortex cluster.:
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.05e-02
Mean rank of genes in gene set: 2091.75
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| NPM1 | 0.0039338 | 109 | GTEx | DepMap | Descartes | 14.22 | 2490.43 |
| MIF | 0.0036348 | 124 | GTEx | DepMap | Descartes | 5.47 | 1671.47 |
| TPI1 | 0.0031444 | 162 | GTEx | DepMap | Descartes | 6.12 | 1000.67 |
| TK1 | -0.0001805 | 7972 | GTEx | DepMap | Descartes | 0.44 | 66.32 |
SCPs mouse unique (Olsen)
Selected list of SCP marker genes shown in Fig. 2B of Olsen et al. https://www.biorxiv.org/content/10.1101/2020.05.04.077057v1 - these genes were unique to mouse SCPs as determined by the linearge tracing study Furlan et al. (Science 2017, PMID 28684471).:
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.63e-02
Mean rank of genes in gene set: 2403
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| H1FX | 0.0037761 | 117 | GTEx | DepMap | Descartes | 7.71 | NA |
| CRABP1 | 0.0018089 | 404 | GTEx | DepMap | Descartes | 0.26 | 111.13 |
| SCRG1 | 0.0002395 | 3326 | GTEx | DepMap | Descartes | 0.06 | 2.47 |
| POSTN | -0.0000385 | 5765 | GTEx | DepMap | Descartes | 0.02 | 2.12 |
Bridge region SCP-adrenergic transition (Olsen)
As above but for cells in the mesenchymal transitioning to SCP-adrenergic region:
Wilcoxon ranksum test P-value for gene set overrepresentation: 2.21e-02
Mean rank of genes in gene set: 2631.75
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| SRP14 | 0.0029926 | 177 | GTEx | DepMap | Descartes | 8.68 | 2295.33 |
| HMGB1 | 0.0015976 | 488 | GTEx | DepMap | Descartes | 16.57 | 836.45 |
| PCBP2 | 0.0000831 | 4450 | GTEx | DepMap | Descartes | 3.66 | 294.33 |
| MDK | -0.0000134 | 5412 | GTEx | DepMap | Descartes | 7.51 | 1585.19 |
Below shows ranks on this GEP for literature curated gene lists for large gene sets
These include those reported as mesenchymal/adrenergic by Van Groningen et al.
High ranks indicate this gene is a driver of this GEP (note these results are not ordered).
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
VanGroningen Adrenergic Genes
Adrenergic marker genes from Supplementary Table 2 of Van Groningen et al. Nature Genetics 2017. These genes were identified by differential expression analysis of mesenchymal-like and adrenergic-like neuroblastoma cell lines.
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.54e-01
Mean rank of genes in gene set: 6619.45
Median rank of genes in gene set: 7005
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| DACH1 | 0.0173399 | 6 | GTEx | DepMap | Descartes | 1.00 | 78.52 |
| HES6 | 0.0130626 | 12 | GTEx | DepMap | Descartes | 1.78 | 502.20 |
| DLK1 | 0.0105537 | 20 | GTEx | DepMap | Descartes | 3.96 | 286.85 |
| NNAT | 0.0069634 | 43 | GTEx | DepMap | Descartes | 3.03 | 975.43 |
| PRSS12 | 0.0051667 | 72 | GTEx | DepMap | Descartes | 0.43 | 31.56 |
| TAGLN3 | 0.0049061 | 77 | GTEx | DepMap | Descartes | 2.53 | 558.62 |
| CKB | 0.0043565 | 85 | GTEx | DepMap | Descartes | 8.25 | 1823.05 |
| SYT1 | 0.0040933 | 102 | GTEx | DepMap | Descartes | 3.27 | 217.65 |
| POPDC3 | 0.0038657 | 112 | GTEx | DepMap | Descartes | 0.32 | 49.83 |
| H1FX | 0.0037761 | 117 | GTEx | DepMap | Descartes | 7.71 | NA |
| VRK1 | 0.0036917 | 122 | GTEx | DepMap | Descartes | 0.55 | 116.03 |
| RANBP1 | 0.0032701 | 148 | GTEx | DepMap | Descartes | 5.52 | 743.99 |
| RFC4 | 0.0032075 | 154 | GTEx | DepMap | Descartes | 0.81 | 195.36 |
| MAP1B | 0.0029991 | 176 | GTEx | DepMap | Descartes | 10.40 | 270.92 |
| ANP32A | 0.0029773 | 178 | GTEx | DepMap | Descartes | 1.75 | 140.19 |
| OLFM1 | 0.0027086 | 210 | GTEx | DepMap | Descartes | 0.61 | 63.99 |
| CEP44 | 0.0026543 | 214 | GTEx | DepMap | Descartes | 0.60 | 44.52 |
| DCX | 0.0024295 | 252 | GTEx | DepMap | Descartes | 2.48 | 70.26 |
| AGTPBP1 | 0.0024071 | 258 | GTEx | DepMap | Descartes | 0.41 | 35.98 |
| HMGA1 | 0.0021470 | 308 | GTEx | DepMap | Descartes | 2.08 | 285.97 |
| MAP6 | 0.0021304 | 311 | GTEx | DepMap | Descartes | 0.70 | 49.43 |
| NEFM | 0.0020750 | 324 | GTEx | DepMap | Descartes | 2.26 | 200.05 |
| NET1 | 0.0019675 | 349 | GTEx | DepMap | Descartes | 0.31 | 24.95 |
| NCS1 | 0.0019537 | 355 | GTEx | DepMap | Descartes | 0.78 | 43.83 |
| TBPL1 | 0.0017703 | 415 | GTEx | DepMap | Descartes | 0.65 | 56.42 |
| UBE2T | 0.0017661 | 417 | GTEx | DepMap | Descartes | 2.03 | 383.23 |
| GAP43 | 0.0017559 | 419 | GTEx | DepMap | Descartes | 2.36 | 355.52 |
| MCM2 | 0.0016435 | 468 | GTEx | DepMap | Descartes | 0.39 | 33.10 |
| TTC8 | 0.0016365 | 471 | GTEx | DepMap | Descartes | 0.26 | 16.44 |
| SYT4 | 0.0014982 | 529 | GTEx | DepMap | Descartes | 0.40 | 29.13 |
VanGroningen Mesenchymal Genes
Mesenchymal marker genes from Supplementary Table 2 of Van Groningen et al. Nature Genetics 2017. These genes were identified by differential expression analysis of mesenchymal-like and adrenergic-like neuroblastoma cell lines.
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 7135.85
Median rank of genes in gene set: 7821
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| SIX1 | 0.0250132 | 2 | GTEx | DepMap | Descartes | 0.65 | 66.49 |
| CYP26A1 | 0.0216206 | 3 | GTEx | DepMap | Descartes | 0.56 | 105.51 |
| ROBO1 | 0.0071305 | 41 | GTEx | DepMap | Descartes | 0.51 | 24.25 |
| FZD7 | 0.0068924 | 45 | GTEx | DepMap | Descartes | 0.22 | 19.51 |
| ADAMTS5 | 0.0058066 | 58 | GTEx | DepMap | Descartes | 0.11 | 5.02 |
| APP | 0.0043473 | 87 | GTEx | DepMap | Descartes | 1.47 | 137.35 |
| PDLIM1 | 0.0040132 | 106 | GTEx | DepMap | Descartes | 0.29 | 75.87 |
| CTNNA1 | 0.0039596 | 107 | GTEx | DepMap | Descartes | 0.60 | 59.84 |
| SSR3 | 0.0036014 | 126 | GTEx | DepMap | Descartes | 1.10 | 102.55 |
| GAS2 | 0.0031737 | 160 | GTEx | DepMap | Descartes | 0.04 | 7.49 |
| SIX4 | 0.0026758 | 213 | GTEx | DepMap | Descartes | 0.04 | 3.52 |
| DUSP14 | 0.0025251 | 239 | GTEx | DepMap | Descartes | 0.15 | 35.06 |
| NR3C1 | 0.0024265 | 253 | GTEx | DepMap | Descartes | 0.27 | 15.98 |
| PHLDB2 | 0.0023303 | 274 | GTEx | DepMap | Descartes | 0.06 | 3.08 |
| PRDX4 | 0.0020386 | 331 | GTEx | DepMap | Descartes | 1.90 | 511.79 |
| ATP1B1 | 0.0018857 | 373 | GTEx | DepMap | Descartes | 1.24 | 141.67 |
| F2R | 0.0018796 | 376 | GTEx | DepMap | Descartes | 0.28 | 22.72 |
| MANF | 0.0018657 | 383 | GTEx | DepMap | Descartes | 1.00 | 211.75 |
| PTPRG | 0.0018434 | 396 | GTEx | DepMap | Descartes | 0.15 | 6.71 |
| WIPI1 | 0.0017718 | 414 | GTEx | DepMap | Descartes | 0.05 | 8.76 |
| PLOD2 | 0.0016960 | 445 | GTEx | DepMap | Descartes | 0.09 | 9.74 |
| TSC22D2 | 0.0016707 | 454 | GTEx | DepMap | Descartes | 0.23 | 6.47 |
| GPC6 | 0.0015750 | 497 | GTEx | DepMap | Descartes | 0.09 | 4.40 |
| RAP1B | 0.0015735 | 498 | GTEx | DepMap | Descartes | 0.73 | 17.07 |
| CKAP4 | 0.0013593 | 637 | GTEx | DepMap | Descartes | 0.56 | 55.71 |
| TUBB6 | 0.0013293 | 654 | GTEx | DepMap | Descartes | 0.11 | 17.52 |
| POLR2L | 0.0013233 | 657 | GTEx | DepMap | Descartes | 1.55 | 553.28 |
| OSTC | 0.0012847 | 688 | GTEx | DepMap | Descartes | 1.26 | 345.51 |
| SERPINE2 | 0.0012695 | 698 | GTEx | DepMap | Descartes | 0.47 | 32.45 |
| ID1 | 0.0012278 | 731 | GTEx | DepMap | Descartes | 0.28 | 83.35 |
Descartes adrenocortical markers
Top 50 marker genes of adrenocortical cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/adrenocortical/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.47e-01
Mean rank of genes in gene set: 7250.39
Median rank of genes in gene set: 7595
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| FDX1 | 0.0012605 | 705 | GTEx | DepMap | Descartes | 0.51 | 46.72 |
| FDPS | 0.0008957 | 1129 | GTEx | DepMap | Descartes | 1.19 | 179.30 |
| CYB5B | 0.0008726 | 1163 | GTEx | DepMap | Descartes | 0.33 | 27.47 |
| MSMO1 | 0.0005092 | 2020 | GTEx | DepMap | Descartes | 0.22 | 28.28 |
| DHCR7 | 0.0004308 | 2320 | GTEx | DepMap | Descartes | 0.09 | 12.76 |
| ERN1 | 0.0001260 | 4080 | GTEx | DepMap | Descartes | 0.04 | 1.81 |
| SLC16A9 | 0.0000968 | 4332 | GTEx | DepMap | Descartes | 0.08 | 4.01 |
| JAKMIP2 | 0.0000340 | 4873 | GTEx | DepMap | Descartes | 0.34 | 11.68 |
| FDXR | 0.0000194 | 5039 | GTEx | DepMap | Descartes | 0.18 | 21.44 |
| TM7SF2 | -0.0000098 | 5365 | GTEx | DepMap | Descartes | 0.33 | 43.69 |
| SCARB1 | -0.0000169 | 5465 | GTEx | DepMap | Descartes | 0.07 | 3.64 |
| SCAP | -0.0000558 | 6014 | GTEx | DepMap | Descartes | 0.19 | 12.89 |
| NPC1 | -0.0000833 | 6443 | GTEx | DepMap | Descartes | 0.03 | 1.58 |
| FREM2 | -0.0000945 | 6615 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| PEG3 | -0.0001056 | 6795 | GTEx | DepMap | Descartes | 0.18 | NA |
| HMGCS1 | -0.0001349 | 7264 | GTEx | DepMap | Descartes | 0.15 | 8.60 |
| PAPSS2 | -0.0001391 | 7323 | GTEx | DepMap | Descartes | 0.01 | 1.40 |
| LDLR | -0.0001496 | 7479 | GTEx | DepMap | Descartes | 0.04 | 2.46 |
| STAR | -0.0001637 | 7711 | GTEx | DepMap | Descartes | 0.00 | 0.18 |
| DHCR24 | -0.0001763 | 7880 | GTEx | DepMap | Descartes | 0.05 | 1.91 |
| PDE10A | -0.0001879 | 8067 | GTEx | DepMap | Descartes | 0.07 | 2.09 |
| HMGCR | -0.0001949 | 8171 | GTEx | DepMap | Descartes | 0.20 | 12.82 |
| SGCZ | -0.0001961 | 8189 | GTEx | DepMap | Descartes | 0.01 | 0.16 |
| FRMD5 | -0.0002297 | 8642 | GTEx | DepMap | Descartes | 0.02 | 1.16 |
| GSTA4 | -0.0002597 | 9007 | GTEx | DepMap | Descartes | 0.78 | 120.17 |
| BAIAP2L1 | -0.0002665 | 9103 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| INHA | -0.0002699 | 9131 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| GRAMD1B | -0.0002841 | 9297 | GTEx | DepMap | Descartes | 0.04 | 1.15 |
| SH3PXD2B | -0.0003360 | 9847 | GTEx | DepMap | Descartes | 0.03 | 1.10 |
| APOC1 | -0.0003513 | 9965 | GTEx | DepMap | Descartes | 2.13 | 607.45 |
Descartes chromaffin markers
Top 50 marker genes of chromaffin cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/chromaffin/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.13e-01
Mean rank of genes in gene set: 7042.46
Median rank of genes in gene set: 8749
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| RYR2 | 0.0037008 | 121 | GTEx | DepMap | Descartes | 0.21 | 4.94 |
| BASP1 | 0.0036703 | 123 | GTEx | DepMap | Descartes | 11.86 | 2029.65 |
| MAB21L2 | 0.0035598 | 134 | GTEx | DepMap | Descartes | 0.68 | 84.53 |
| KCNB2 | 0.0033630 | 144 | GTEx | DepMap | Descartes | 0.20 | 22.37 |
| MAP1B | 0.0029991 | 176 | GTEx | DepMap | Descartes | 10.40 | 270.92 |
| TUBA1A | 0.0029652 | 179 | GTEx | DepMap | Descartes | 31.25 | 5294.46 |
| GAP43 | 0.0017559 | 419 | GTEx | DepMap | Descartes | 2.36 | 355.52 |
| ANKFN1 | 0.0012915 | 681 | GTEx | DepMap | Descartes | 0.05 | 3.72 |
| MLLT11 | 0.0010498 | 914 | GTEx | DepMap | Descartes | 5.11 | 582.81 |
| HS3ST5 | 0.0009732 | 1008 | GTEx | DepMap | Descartes | 0.05 | 5.01 |
| ELAVL2 | 0.0009413 | 1048 | GTEx | DepMap | Descartes | 0.58 | 45.43 |
| EYA1 | 0.0009218 | 1079 | GTEx | DepMap | Descartes | 0.12 | 10.92 |
| TUBB2B | 0.0003923 | 2495 | GTEx | DepMap | Descartes | 13.15 | 2164.28 |
| SYNPO2 | 0.0003847 | 2527 | GTEx | DepMap | Descartes | 0.26 | 4.26 |
| EPHA6 | 0.0002771 | 3080 | GTEx | DepMap | Descartes | 0.02 | 2.55 |
| GAL | 0.0001968 | 3572 | GTEx | DepMap | Descartes | 1.92 | 666.41 |
| RBFOX1 | 0.0001255 | 4088 | GTEx | DepMap | Descartes | 0.18 | 10.79 |
| TUBB2A | 0.0000542 | 4701 | GTEx | DepMap | Descartes | 2.06 | 352.77 |
| SLC44A5 | -0.0001242 | 7092 | GTEx | DepMap | Descartes | 0.06 | 3.53 |
| CNKSR2 | -0.0001662 | 7744 | GTEx | DepMap | Descartes | 0.09 | 2.74 |
| CCND1 | -0.0002376 | 8749 | GTEx | DepMap | Descartes | 7.20 | 474.38 |
| GREM1 | -0.0004312 | 10575 | GTEx | DepMap | Descartes | 0.02 | 0.70 |
| STMN2 | -0.0004712 | 10867 | GTEx | DepMap | Descartes | 16.16 | 2219.68 |
| RPH3A | -0.0005560 | 11297 | GTEx | DepMap | Descartes | 0.09 | 3.83 |
| REEP1 | -0.0006160 | 11516 | GTEx | DepMap | Descartes | 0.23 | 12.07 |
| IL7 | -0.0006556 | 11639 | GTEx | DepMap | Descartes | 0.01 | 1.02 |
| PTCHD1 | -0.0006632 | 11664 | GTEx | DepMap | Descartes | 0.03 | 0.76 |
| RGMB | -0.0006679 | 11681 | GTEx | DepMap | Descartes | 0.22 | 11.11 |
| NTRK1 | -0.0006689 | 11687 | GTEx | DepMap | Descartes | 0.08 | 9.66 |
| PLXNA4 | -0.0008055 | 11982 | GTEx | DepMap | Descartes | 0.06 | 0.95 |
Descartes Vascular_endothelial markers
Top 50 marker genes of Vascular_endothelial cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/vascular_endothelial/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.14e-01
Mean rank of genes in gene set: 7074.63
Median rank of genes in gene set: 6906.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CDH13 | 0.0041373 | 99 | GTEx | DepMap | Descartes | 0.12 | 6.51 |
| CLDN5 | 0.0018592 | 387 | GTEx | DepMap | Descartes | 0.22 | 37.28 |
| ID1 | 0.0012278 | 731 | GTEx | DepMap | Descartes | 0.28 | 83.35 |
| CALCRL | 0.0005317 | 1943 | GTEx | DepMap | Descartes | 0.01 | 1.36 |
| RASIP1 | 0.0003634 | 2641 | GTEx | DepMap | Descartes | 0.02 | 2.05 |
| NPR1 | 0.0003041 | 2935 | GTEx | DepMap | Descartes | 0.01 | 0.55 |
| CRHBP | -0.0000237 | 5558 | GTEx | DepMap | Descartes | 0.00 | 0.37 |
| NOTCH4 | -0.0000438 | 5850 | GTEx | DepMap | Descartes | 0.10 | 3.32 |
| CDH5 | -0.0000440 | 5854 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| F8 | -0.0000601 | 6083 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| KANK3 | -0.0000623 | 6117 | GTEx | DepMap | Descartes | 0.00 | 0.35 |
| SHANK3 | -0.0000780 | 6355 | GTEx | DepMap | Descartes | 0.01 | 0.43 |
| FLT4 | -0.0000790 | 6369 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ESM1 | -0.0000828 | 6429 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SLCO2A1 | -0.0000994 | 6692 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| GALNT15 | -0.0001002 | 6711 | GTEx | DepMap | Descartes | 0.00 | NA |
| TIE1 | -0.0001054 | 6789 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ROBO4 | -0.0001058 | 6801 | GTEx | DepMap | Descartes | 0.00 | 0.13 |
| KDR | -0.0001061 | 6808 | GTEx | DepMap | Descartes | 0.00 | 0.10 |
| MYRIP | -0.0001181 | 7005 | GTEx | DepMap | Descartes | 0.02 | 1.35 |
| SHE | -0.0001306 | 7207 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| MMRN2 | -0.0001351 | 7267 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| CEACAM1 | -0.0001522 | 7513 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| BTNL9 | -0.0001585 | 7623 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| PLVAP | -0.0001685 | 7775 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| NR5A2 | -0.0001777 | 7915 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| PODXL | -0.0001941 | 8158 | GTEx | DepMap | Descartes | 0.03 | 0.74 |
| HYAL2 | -0.0002194 | 8518 | GTEx | DepMap | Descartes | 0.43 | 32.95 |
| TEK | -0.0002321 | 8673 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| EHD3 | -0.0002637 | 9075 | GTEx | DepMap | Descartes | 0.01 | 0.60 |
Descartes stromal markers
Top 50 marker genes of stromal cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/stromal/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.74e-01
Mean rank of genes in gene set: 7334.09
Median rank of genes in gene set: 7935.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| GAS2 | 0.0031737 | 160 | GTEx | DepMap | Descartes | 0.04 | 7.49 |
| LOX | 0.0016144 | 479 | GTEx | DepMap | Descartes | 0.05 | 4.91 |
| HHIP | 0.0012991 | 670 | GTEx | DepMap | Descartes | 0.41 | 13.30 |
| ABCC9 | 0.0008453 | 1220 | GTEx | DepMap | Descartes | 0.01 | 0.82 |
| SFRP2 | 0.0000977 | 4327 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| PCDH18 | 0.0000926 | 4365 | GTEx | DepMap | Descartes | 0.01 | 0.40 |
| FREM1 | 0.0000786 | 4490 | GTEx | DepMap | Descartes | 0.00 | 0.04 |
| ITGA11 | 0.0000286 | 4940 | GTEx | DepMap | Descartes | 0.00 | 0.13 |
| POSTN | -0.0000385 | 5765 | GTEx | DepMap | Descartes | 0.02 | 2.12 |
| RSPO3 | -0.0000486 | 5923 | GTEx | DepMap | Descartes | 0.00 | NA |
| COL6A3 | -0.0000578 | 6049 | GTEx | DepMap | Descartes | 0.02 | 0.42 |
| GLI2 | -0.0000633 | 6139 | GTEx | DepMap | Descartes | 0.00 | 0.20 |
| BICC1 | -0.0000710 | 6246 | GTEx | DepMap | Descartes | 0.01 | 0.87 |
| CCDC80 | -0.0000856 | 6479 | GTEx | DepMap | Descartes | 0.02 | 0.60 |
| SCARA5 | -0.0001094 | 6862 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ADAMTSL3 | -0.0001099 | 6870 | GTEx | DepMap | Descartes | 0.00 | 0.02 |
| CLDN11 | -0.0001159 | 6973 | GTEx | DepMap | Descartes | 0.02 | 0.43 |
| CD248 | -0.0001471 | 7443 | GTEx | DepMap | Descartes | 0.02 | 1.27 |
| PAMR1 | -0.0001518 | 7509 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ACTA2 | -0.0001632 | 7700 | GTEx | DepMap | Descartes | 0.07 | 11.61 |
| OGN | -0.0001693 | 7785 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| DCN | -0.0001782 | 7925 | GTEx | DepMap | Descartes | 0.00 | 0.09 |
| EDNRA | -0.0001791 | 7946 | GTEx | DepMap | Descartes | 0.00 | 0.04 |
| IGFBP3 | -0.0001794 | 7951 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ABCA6 | -0.0001803 | 7968 | GTEx | DepMap | Descartes | 0.00 | 0.11 |
| LUM | -0.0001829 | 8007 | GTEx | DepMap | Descartes | 0.02 | 1.39 |
| COL27A1 | -0.0001848 | 8032 | GTEx | DepMap | Descartes | 0.00 | 0.06 |
| LAMC3 | -0.0001891 | 8080 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| MGP | -0.0001922 | 8139 | GTEx | DepMap | Descartes | 0.04 | 1.43 |
| DKK2 | -0.0001947 | 8166 | GTEx | DepMap | Descartes | 0.00 | 0.25 |
Descartes sympathoblasts markers
Top 50 marker genes of sympathoblasts cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/sympathoblasts/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.95e-01
Mean rank of genes in gene set: 7779.55
Median rank of genes in gene set: 8155
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| ST18 | 0.0114264 | 17 | GTEx | DepMap | Descartes | 0.14 | 9.51 |
| ROBO1 | 0.0071305 | 41 | GTEx | DepMap | Descartes | 0.51 | 24.25 |
| CCSER1 | 0.0018774 | 379 | GTEx | DepMap | Descartes | 0.09 | NA |
| GCH1 | 0.0006865 | 1518 | GTEx | DepMap | Descartes | 0.12 | 12.15 |
| HTATSF1 | 0.0005071 | 2031 | GTEx | DepMap | Descartes | 0.78 | 73.56 |
| ARC | 0.0004578 | 2215 | GTEx | DepMap | Descartes | 0.86 | 97.46 |
| SLC24A2 | 0.0003256 | 2813 | GTEx | DepMap | Descartes | 0.01 | 0.25 |
| CNTN3 | 0.0000722 | 4544 | GTEx | DepMap | Descartes | 0.00 | 0.07 |
| PENK | 0.0000392 | 4832 | GTEx | DepMap | Descartes | 0.00 | 0.79 |
| SLC35F3 | 0.0000267 | 4957 | GTEx | DepMap | Descartes | 0.01 | 0.23 |
| CDH12 | -0.0000517 | 5962 | GTEx | DepMap | Descartes | 0.05 | 3.02 |
| TENM1 | -0.0000566 | 6027 | GTEx | DepMap | Descartes | 0.01 | NA |
| CHGA | -0.0000619 | 6109 | GTEx | DepMap | Descartes | 1.29 | 197.45 |
| AGBL4 | -0.0000864 | 6493 | GTEx | DepMap | Descartes | 0.02 | 0.48 |
| GALNTL6 | -0.0001140 | 6944 | GTEx | DepMap | Descartes | 0.03 | 3.86 |
| GRM7 | -0.0001285 | 7166 | GTEx | DepMap | Descartes | 0.00 | 0.08 |
| TIAM1 | -0.0001532 | 7534 | GTEx | DepMap | Descartes | 0.16 | 6.13 |
| PCSK2 | -0.0001764 | 7881 | GTEx | DepMap | Descartes | 0.00 | 0.13 |
| KSR2 | -0.0001885 | 8074 | GTEx | DepMap | Descartes | 0.05 | 0.66 |
| SPOCK3 | -0.0001996 | 8236 | GTEx | DepMap | Descartes | 0.02 | 1.16 |
| EML6 | -0.0002009 | 8260 | GTEx | DepMap | Descartes | 0.01 | 0.26 |
| LAMA3 | -0.0002301 | 8650 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| CDH18 | -0.0002335 | 8695 | GTEx | DepMap | Descartes | 0.00 | 0.12 |
| TBX20 | -0.0003121 | 9599 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SORCS3 | -0.0004540 | 10760 | GTEx | DepMap | Descartes | 0.08 | 3.96 |
| DGKK | -0.0005585 | 11309 | GTEx | DepMap | Descartes | 0.04 | 1.21 |
| SLC18A1 | -0.0005834 | 11400 | GTEx | DepMap | Descartes | 0.00 | 0.24 |
| MGAT4C | -0.0005902 | 11428 | GTEx | DepMap | Descartes | 0.04 | 0.26 |
| FAM155A | -0.0006968 | 11744 | GTEx | DepMap | Descartes | 0.24 | 6.29 |
| CHGB | -0.0007866 | 11948 | GTEx | DepMap | Descartes | 0.85 | 99.03 |
Descartes erythroblasts markers
Top 50 marker genes of erythroblasts cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/erythroblasts/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.77e-01
Mean rank of genes in gene set: 7615.1
Median rank of genes in gene set: 6954
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| SLC25A37 | 0.0002633 | 3163 | GTEx | DepMap | Descartes | 0.29 | 17.73 |
| ANK1 | 0.0001999 | 3555 | GTEx | DepMap | Descartes | 0.04 | 1.58 |
| BLVRB | 0.0001646 | 3774 | GTEx | DepMap | Descartes | 0.27 | 53.27 |
| SLC25A21 | 0.0001639 | 3780 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| CPOX | 0.0000396 | 4829 | GTEx | DepMap | Descartes | 0.05 | 6.15 |
| XPO7 | 0.0000113 | 5134 | GTEx | DepMap | Descartes | 0.15 | 9.13 |
| RGS6 | 0.0000049 | 5195 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| FECH | -0.0000018 | 5274 | GTEx | DepMap | Descartes | 0.07 | 2.67 |
| SNCA | -0.0000060 | 5326 | GTEx | DepMap | Descartes | 0.20 | 12.08 |
| RHD | -0.0000660 | 6181 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| TMCC2 | -0.0000764 | 6324 | GTEx | DepMap | Descartes | 0.02 | 1.15 |
| SLC4A1 | -0.0000771 | 6342 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ALAS2 | -0.0000923 | 6582 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| TRAK2 | -0.0000993 | 6689 | GTEx | DepMap | Descartes | 0.09 | 3.74 |
| SELENBP1 | -0.0001145 | 6954 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ABCB10 | -0.0001198 | 7029 | GTEx | DepMap | Descartes | 0.02 | 1.21 |
| MARCH3 | -0.0001776 | 7913 | GTEx | DepMap | Descartes | 0.05 | NA |
| GCLC | -0.0001849 | 8035 | GTEx | DepMap | Descartes | 0.07 | 5.22 |
| DENND4A | -0.0001970 | 8208 | GTEx | DepMap | Descartes | 0.14 | 5.15 |
| MICAL2 | -0.0002459 | 8858 | GTEx | DepMap | Descartes | 0.01 | 0.69 |
| TFR2 | -0.0002832 | 9289 | GTEx | DepMap | Descartes | 0.06 | 5.32 |
| SPTB | -0.0004673 | 10836 | GTEx | DepMap | Descartes | 0.04 | 0.55 |
| SPECC1 | -0.0005231 | 11163 | GTEx | DepMap | Descartes | 0.06 | 2.36 |
| SOX6 | -0.0005395 | 11230 | GTEx | DepMap | Descartes | 0.01 | 0.12 |
| EPB41 | -0.0005697 | 11358 | GTEx | DepMap | Descartes | 0.24 | 11.36 |
| GYPC | -0.0006103 | 11495 | GTEx | DepMap | Descartes | 0.05 | 3.57 |
| CAT | -0.0007624 | 11909 | GTEx | DepMap | Descartes | 0.13 | 13.46 |
| RAPGEF2 | -0.0008578 | 12068 | GTEx | DepMap | Descartes | 0.17 | 5.31 |
| TSPAN5 | -0.0011589 | 12345 | GTEx | DepMap | Descartes | 0.28 | 15.85 |
| NA | NA | NA | NA | NA | NA | NA | NA |
Descartes myeloid markers
Top 50 marker genes of myeloid cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/myeloid/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.96e-01
Mean rank of genes in gene set: 7844.34
Median rank of genes in gene set: 8927
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| RBPJ | 0.0010417 | 925 | GTEx | DepMap | Descartes | 0.97 | 45.98 |
| MARCH1 | 0.0010334 | 937 | GTEx | DepMap | Descartes | 0.09 | NA |
| CTSC | 0.0009453 | 1044 | GTEx | DepMap | Descartes | 0.41 | 28.41 |
| CD163L1 | 0.0008638 | 1183 | GTEx | DepMap | Descartes | 0.15 | 13.48 |
| SFMBT2 | 0.0004747 | 2146 | GTEx | DepMap | Descartes | 0.06 | 2.14 |
| HCK | 0.0000545 | 4700 | GTEx | DepMap | Descartes | 0.01 | 1.21 |
| FMN1 | 0.0000287 | 4939 | GTEx | DepMap | Descartes | 0.07 | 0.66 |
| ATP8B4 | -0.0000345 | 5704 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| FGD2 | -0.0000841 | 6454 | GTEx | DepMap | Descartes | 0.01 | 0.27 |
| MERTK | -0.0001027 | 6746 | GTEx | DepMap | Descartes | 0.02 | 1.11 |
| AXL | -0.0001033 | 6761 | GTEx | DepMap | Descartes | 0.03 | 1.18 |
| ITPR2 | -0.0001313 | 7216 | GTEx | DepMap | Descartes | 0.20 | 4.69 |
| IFNGR1 | -0.0001361 | 7284 | GTEx | DepMap | Descartes | 0.18 | 21.26 |
| CST3 | -0.0001494 | 7474 | GTEx | DepMap | Descartes | 1.77 | 145.96 |
| LGMN | -0.0001618 | 7678 | GTEx | DepMap | Descartes | 0.34 | 32.93 |
| SLC1A3 | -0.0001809 | 7974 | GTEx | DepMap | Descartes | 0.00 | 0.15 |
| CPVL | -0.0001986 | 8226 | GTEx | DepMap | Descartes | 0.04 | 4.59 |
| HRH1 | -0.0002244 | 8579 | GTEx | DepMap | Descartes | 0.00 | 0.08 |
| SPP1 | -0.0002345 | 8718 | GTEx | DepMap | Descartes | 13.63 | 1642.60 |
| FGL2 | -0.0002704 | 9136 | GTEx | DepMap | Descartes | 0.02 | 1.21 |
| MSR1 | -0.0002718 | 9149 | GTEx | DepMap | Descartes | 0.05 | 2.27 |
| PTPRE | -0.0002799 | 9253 | GTEx | DepMap | Descartes | 0.03 | 0.91 |
| CD163 | -0.0002980 | 9453 | GTEx | DepMap | Descartes | 0.07 | 2.15 |
| MS4A4A | -0.0003223 | 9708 | GTEx | DepMap | Descartes | 0.07 | 6.81 |
| CYBB | -0.0003537 | 9985 | GTEx | DepMap | Descartes | 0.01 | 0.23 |
| SLC9A9 | -0.0003585 | 10024 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SLCO2B1 | -0.0003603 | 10045 | GTEx | DepMap | Descartes | 0.02 | 0.52 |
| RGL1 | -0.0003686 | 10132 | GTEx | DepMap | Descartes | 0.03 | 1.33 |
| CD14 | -0.0003790 | 10210 | GTEx | DepMap | Descartes | 0.36 | 41.61 |
| CSF1R | -0.0004032 | 10382 | GTEx | DepMap | Descartes | 0.02 | 0.81 |
Descartes Schwann markers
Top 50 marker genes of Schwann cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/schwann/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.94e-01
Mean rank of genes in gene set: 7655.93
Median rank of genes in gene set: 8205
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| ADAMTS5 | 0.0058066 | 58 | GTEx | DepMap | Descartes | 0.11 | 5.02 |
| EGFLAM | 0.0055140 | 64 | GTEx | DepMap | Descartes | 0.21 | 19.54 |
| FIGN | 0.0031978 | 155 | GTEx | DepMap | Descartes | 0.34 | 10.83 |
| MARCKS | 0.0012066 | 759 | GTEx | DepMap | Descartes | 7.18 | 468.80 |
| XKR4 | 0.0006367 | 1647 | GTEx | DepMap | Descartes | 0.02 | 0.19 |
| HMGA2 | 0.0004806 | 2129 | GTEx | DepMap | Descartes | 0.00 | 0.13 |
| NRXN3 | 0.0004466 | 2258 | GTEx | DepMap | Descartes | 0.02 | 0.58 |
| LRRTM4 | 0.0003673 | 2624 | GTEx | DepMap | Descartes | 0.02 | 3.26 |
| SLC35F1 | 0.0002665 | 3140 | GTEx | DepMap | Descartes | 0.08 | 3.37 |
| PTPRZ1 | 0.0001093 | 4228 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| KCTD12 | 0.0000655 | 4607 | GTEx | DepMap | Descartes | 0.07 | 3.26 |
| SFRP1 | 0.0000306 | 4918 | GTEx | DepMap | Descartes | 0.22 | 18.00 |
| IL1RAPL2 | -0.0000093 | 5361 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ERBB3 | -0.0000215 | 5518 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| OLFML2A | -0.0000425 | 5824 | GTEx | DepMap | Descartes | 0.00 | 0.03 |
| PLP1 | -0.0000665 | 6187 | GTEx | DepMap | Descartes | 0.01 | 0.75 |
| MDGA2 | -0.0000801 | 6384 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| GRIK3 | -0.0001090 | 6850 | GTEx | DepMap | Descartes | 0.00 | 0.25 |
| MPZ | -0.0001368 | 7295 | GTEx | DepMap | Descartes | 0.00 | 0.23 |
| EDNRB | -0.0001538 | 7543 | GTEx | DepMap | Descartes | 0.01 | 0.19 |
| TRPM3 | -0.0001594 | 7637 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| COL18A1 | -0.0001846 | 8031 | GTEx | DepMap | Descartes | 0.03 | 0.62 |
| SORCS1 | -0.0002093 | 8379 | GTEx | DepMap | Descartes | 0.03 | 0.79 |
| COL5A2 | -0.0002209 | 8536 | GTEx | DepMap | Descartes | 0.04 | 0.86 |
| STARD13 | -0.0002485 | 8886 | GTEx | DepMap | Descartes | 0.01 | 0.20 |
| COL25A1 | -0.0003144 | 9625 | GTEx | DepMap | Descartes | 0.02 | 0.50 |
| LAMC1 | -0.0003438 | 9903 | GTEx | DepMap | Descartes | 0.03 | 1.06 |
| NRXN1 | -0.0003470 | 9929 | GTEx | DepMap | Descartes | 0.85 | 22.75 |
| GAS7 | -0.0003778 | 10200 | GTEx | DepMap | Descartes | 0.00 | 0.03 |
| IL1RAPL1 | -0.0004573 | 10777 | GTEx | DepMap | Descartes | 0.00 | 0.07 |
Descartes Megakaryocytes markers
Top 50 marker genes of Megakaryocytes cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/megakaryocytes/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.85e-01
Mean rank of genes in gene set: 7445.11
Median rank of genes in gene set: 8359
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| TMSB4X | 0.0024224 | 254 | GTEx | DepMap | Descartes | 31.22 | 6610.58 |
| RAP1B | 0.0015735 | 498 | GTEx | DepMap | Descartes | 0.73 | 17.07 |
| MED12L | 0.0007457 | 1407 | GTEx | DepMap | Descartes | 0.04 | 1.12 |
| TPM4 | 0.0006499 | 1618 | GTEx | DepMap | Descartes | 1.27 | 78.14 |
| LIMS1 | 0.0004342 | 2302 | GTEx | DepMap | Descartes | 0.50 | 33.26 |
| PSTPIP2 | 0.0003492 | 2711 | GTEx | DepMap | Descartes | 0.02 | 2.49 |
| ANGPT1 | 0.0003077 | 2916 | GTEx | DepMap | Descartes | 0.03 | 1.20 |
| SLC24A3 | 0.0001543 | 3850 | GTEx | DepMap | Descartes | 0.01 | 0.68 |
| CD9 | 0.0000931 | 4362 | GTEx | DepMap | Descartes | 0.44 | 89.41 |
| VCL | 0.0000731 | 4534 | GTEx | DepMap | Descartes | 0.12 | 5.08 |
| ITGB3 | 0.0000570 | 4682 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ITGA2B | 0.0000293 | 4934 | GTEx | DepMap | Descartes | 0.03 | 2.63 |
| FLNA | 0.0000243 | 4978 | GTEx | DepMap | Descartes | 0.23 | 8.29 |
| STON2 | -0.0000082 | 5348 | GTEx | DepMap | Descartes | 0.04 | 2.61 |
| ARHGAP6 | -0.0000250 | 5582 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| P2RX1 | -0.0000806 | 6392 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| MMRN1 | -0.0000905 | 6553 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| TUBB1 | -0.0000998 | 6701 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| RAB27B | -0.0000999 | 6706 | GTEx | DepMap | Descartes | 0.02 | 0.40 |
| GP1BA | -0.0001043 | 6773 | GTEx | DepMap | Descartes | 0.00 | 0.07 |
| LTBP1 | -0.0001071 | 6822 | GTEx | DepMap | Descartes | 0.02 | 0.62 |
| STOM | -0.0001254 | 7112 | GTEx | DepMap | Descartes | 0.08 | 6.56 |
| UBASH3B | -0.0002075 | 8359 | GTEx | DepMap | Descartes | 0.03 | 0.77 |
| THBS1 | -0.0002269 | 8614 | GTEx | DepMap | Descartes | 0.00 | 0.17 |
| PLEK | -0.0002299 | 8646 | GTEx | DepMap | Descartes | 0.09 | 6.34 |
| TRPC6 | -0.0002312 | 8666 | GTEx | DepMap | Descartes | 0.00 | 0.04 |
| GSN | -0.0002446 | 8845 | GTEx | DepMap | Descartes | 0.04 | 1.58 |
| FLI1 | -0.0002455 | 8853 | GTEx | DepMap | Descartes | 0.01 | 0.42 |
| MCTP1 | -0.0002812 | 9267 | GTEx | DepMap | Descartes | 0.00 | 0.07 |
| MYLK | -0.0002927 | 9402 | GTEx | DepMap | Descartes | 0.02 | 0.15 |
Descartes Lyphoid markers
Top 50 marker genes of Lyphoid cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/lymphoid/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 9878.05
Median rank of genes in gene set: 10749.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| TMSB10 | 0.0017544 | 421 | GTEx | DepMap | Descartes | 46.91 | 28707.38 |
| FYN | 0.0006656 | 1571 | GTEx | DepMap | Descartes | 0.91 | 74.29 |
| NCALD | 0.0005440 | 1909 | GTEx | DepMap | Descartes | 0.06 | 2.74 |
| MBNL1 | 0.0000954 | 4345 | GTEx | DepMap | Descartes | 0.18 | 7.16 |
| CCND3 | 0.0000850 | 4429 | GTEx | DepMap | Descartes | 0.15 | 17.10 |
| PDE3B | -0.0002096 | 8384 | GTEx | DepMap | Descartes | 0.10 | 4.04 |
| SAMD3 | -0.0002271 | 8616 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ITPKB | -0.0002360 | 8737 | GTEx | DepMap | Descartes | 0.00 | 0.25 |
| CCL5 | -0.0002460 | 8862 | GTEx | DepMap | Descartes | 0.04 | 11.46 |
| ANKRD44 | -0.0002554 | 8966 | GTEx | DepMap | Descartes | 0.08 | 2.61 |
| SKAP1 | -0.0002788 | 9237 | GTEx | DepMap | Descartes | 0.01 | 3.23 |
| RCSD1 | -0.0003234 | 9720 | GTEx | DepMap | Descartes | 0.00 | 0.10 |
| ARHGAP15 | -0.0003254 | 9743 | GTEx | DepMap | Descartes | 0.00 | 0.20 |
| PRKCH | -0.0003269 | 9758 | GTEx | DepMap | Descartes | 0.01 | 0.37 |
| MCTP2 | -0.0003358 | 9842 | GTEx | DepMap | Descartes | 0.01 | 0.18 |
| PLEKHA2 | -0.0003553 | 9999 | GTEx | DepMap | Descartes | 0.02 | 0.59 |
| IKZF1 | -0.0003844 | 10246 | GTEx | DepMap | Descartes | 0.01 | 0.64 |
| LEF1 | -0.0004214 | 10501 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| EVL | -0.0004388 | 10636 | GTEx | DepMap | Descartes | 1.65 | 123.86 |
| SP100 | -0.0004453 | 10688 | GTEx | DepMap | Descartes | 0.01 | 0.31 |
| WIPF1 | -0.0004518 | 10740 | GTEx | DepMap | Descartes | 0.03 | 1.63 |
| DOCK10 | -0.0004540 | 10759 | GTEx | DepMap | Descartes | 0.03 | 0.92 |
| SCML4 | -0.0005010 | 11037 | GTEx | DepMap | Descartes | 0.01 | 0.22 |
| ETS1 | -0.0005087 | 11084 | GTEx | DepMap | Descartes | 0.01 | 0.23 |
| PTPRC | -0.0005515 | 11279 | GTEx | DepMap | Descartes | 0.02 | 0.75 |
| STK39 | -0.0005666 | 11341 | GTEx | DepMap | Descartes | 0.14 | 8.40 |
| GNG2 | -0.0005818 | 11395 | GTEx | DepMap | Descartes | 0.44 | 24.06 |
| LCP1 | -0.0006389 | 11596 | GTEx | DepMap | Descartes | 0.04 | 1.92 |
| ARHGDIB | -0.0006695 | 11690 | GTEx | DepMap | Descartes | 0.13 | 26.38 |
| FOXP1 | -0.0006924 | 11733 | GTEx | DepMap | Descartes | 0.55 | 15.45 |
Below shows the ranks on this GEP for immune marker gene lists curated by CellTypist (https://www.celltypist.org/encyclopedia).
High ranks indicate this gene is a driver of this GEP.
These curated gene list are ranked by P-value (on this GEP) of their constituent genes.
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
ILC precursor: ILC precursor (model markers)
innate lymphoid cell precursors which give rise to innate lymphoid cells and lack the characteristics of their mature progenies:
Wilcoxon ranksum test P-value for gene set overrepresentation: 2.01e-02
Mean rank of genes in gene set: 1985
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| HPN | 0.0007075 | 1474 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| HUNK | 0.0005164 | 1992 | GTEx | DepMap | Descartes | 0.11 | 5.82 |
| PDZK1 | 0.0003937 | 2489 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
HSC/MPP: Early lymphoid/T lymphoid (model markers)
early lymphoid/T lymphocytes with lymphocyte potential in the fetal liver before T cells emerged from the thymus:
Wilcoxon ranksum test P-value for gene set overrepresentation: 4.21e-02
Mean rank of genes in gene set: 17
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| ST18 | 0.0114264 | 17 | GTEx | DepMap | Descartes | 0.14 | 9.51 |
HSC/MPP: Megakaryocyte-erythroid-mast cell progenitor (model markers)
shared progenitors in the fetal liver which originate from common myeloid progenitors and differentiate into megakaryocytes:
Wilcoxon ranksum test P-value for gene set overrepresentation: 4.38e-02
Mean rank of genes in gene set: 85
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CKB | 0.0043565 | 85 | GTEx | DepMap | Descartes | 8.25 | 1823.05 |