Program: 13. Intermediate Monocyte.
Submit a comment on this gene expression program’s interpretation: CLICK
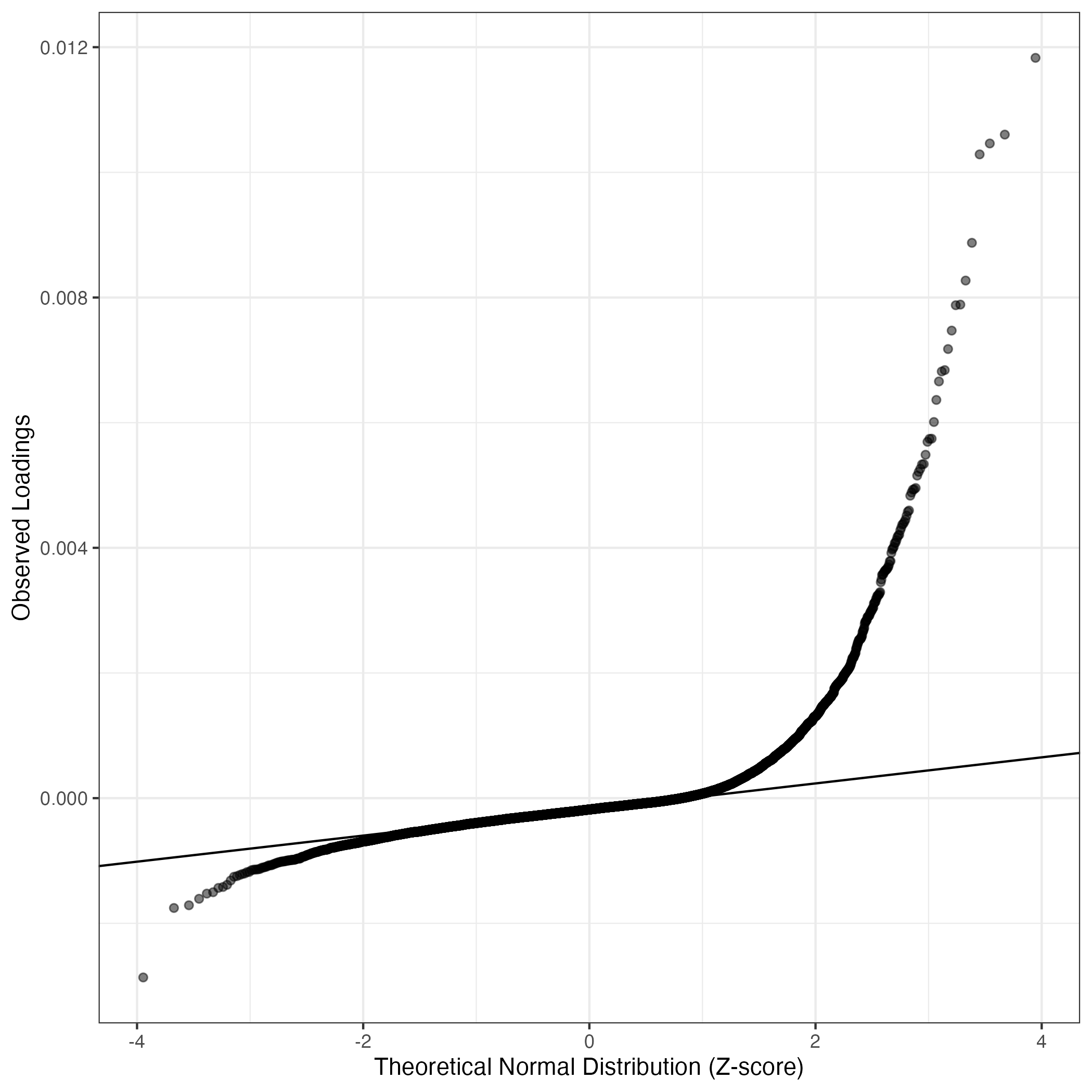
QQ-plot of gene loadings, averaged over both independent splits of the data
This plot highlights the relative contribution of each gene to the GEP
Top genes driving this program. Note: Decartes website is buggy, try refreshing. Also, Decartes fetal adrenal data have been collected at specific time points (89-122 days), all possible cell types of interest may not be represented, do not overinterpret. The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program (currently only calculated for the cells in the first 50/50 split of the data).
Note: Decartes website is buggy, try refreshing. Also, Decartes fetal adrenal data have been collected at specific time points (89-122 days), all possible cell types of interest may not be represented, do not overinterpret.
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
| Gene | Loading | Gene.Name | GTEx | DepMap | Descartes | Mean.Counts | Mean.Tpm | |
|---|---|---|---|---|---|---|---|---|
| 1 | CD74 | 0.0118294 | CD74 molecule | GTEx | DepMap | Descartes | 212.37 | 16090.36 |
| 2 | IDO1 | 0.0106031 | indoleamine 2,3-dioxygenase 1 | GTEx | DepMap | Descartes | 3.06 | 359.51 |
| 3 | LAMP3 | 0.0104613 | lysosomal associated membrane protein 3 | GTEx | DepMap | Descartes | 1.75 | 122.40 |
| 4 | CCR7 | 0.0102872 | C-C motif chemokine receptor 7 | GTEx | DepMap | Descartes | 17.23 | 2204.68 |
| 5 | DAPP1 | 0.0088738 | dual adaptor of phosphotyrosine and 3-phosphoinositides 1 | GTEx | DepMap | Descartes | 1.62 | 130.15 |
| 6 | CD83 | 0.0082708 | CD83 molecule | GTEx | DepMap | Descartes | 14.60 | 1555.01 |
| 7 | CYP2S1 | 0.0078867 | cytochrome P450 family 2 subfamily S member 1 | GTEx | DepMap | Descartes | 0.57 | 59.49 |
| 8 | GPR157 | 0.0078768 | G protein-coupled receptor 157 | GTEx | DepMap | Descartes | 0.91 | 54.72 |
| 9 | SLC7A11 | 0.0074713 | solute carrier family 7 member 11 | GTEx | DepMap | Descartes | 4.76 | 113.38 |
| 10 | LAD1 | 0.0071765 | ladinin 1 | GTEx | DepMap | Descartes | 0.42 | 37.06 |
| 11 | SERPINB9 | 0.0068400 | serpin family B member 9 | GTEx | DepMap | Descartes | 12.84 | 837.95 |
| 12 | CST3 | 0.0068182 | cystatin C | GTEx | DepMap | Descartes | 23.92 | 1682.07 |
| 13 | SLC6A12 | 0.0066590 | solute carrier family 6 member 12 | GTEx | DepMap | Descartes | 0.37 | 16.11 |
| 14 | CPVL | 0.0063634 | carboxypeptidase vitellogenic like | GTEx | DepMap | Descartes | 3.57 | 349.47 |
| 15 | GPR183 | 0.0060107 | G protein-coupled receptor 183 | GTEx | DepMap | Descartes | 13.08 | 2254.53 |
| 16 | OGFRL1 | 0.0057437 | opioid growth factor receptor like 1 | GTEx | DepMap | Descartes | 3.32 | 95.67 |
| 17 | KYNU | 0.0057416 | kynureninase | GTEx | DepMap | Descartes | 4.86 | 80.79 |
| 18 | RASAL1 | 0.0056941 | RAS protein activator like 1 | GTEx | DepMap | Descartes | 0.16 | 9.19 |
| 19 | CMTM6 | 0.0054872 | CKLF like MARVEL transmembrane domain containing 6 | GTEx | DepMap | Descartes | 6.99 | 565.46 |
| 20 | SPI1 | 0.0053386 | Spi-1 proto-oncogene | GTEx | DepMap | Descartes | 3.28 | 556.98 |
| 21 | WDFY4 | 0.0053335 | WDFY family member 4 | GTEx | DepMap | Descartes | 0.68 | 15.56 |
| 22 | G0S2 | 0.0052635 | G0/G1 switch 2 | GTEx | DepMap | Descartes | 14.57 | 4553.91 |
| 23 | LGALS2 | 0.0052135 | galectin 2 | GTEx | DepMap | Descartes | 0.76 | 304.07 |
| 24 | BIRC3 | 0.0051541 | baculoviral IAP repeat containing 3 | GTEx | DepMap | Descartes | 14.47 | 605.61 |
| 25 | SLAMF7 | 0.0049559 | SLAM family member 7 | GTEx | DepMap | Descartes | 4.05 | 276.49 |
| 26 | NR4A3 | 0.0049356 | nuclear receptor subfamily 4 group A member 3 | GTEx | DepMap | Descartes | 5.13 | 236.18 |
| 27 | PADI2 | 0.0049306 | peptidyl arginine deiminase 2 | GTEx | DepMap | Descartes | 0.49 | 38.47 |
| 28 | MIR155HG | 0.0048851 | MIR155 host gene | GTEx | DepMap | Descartes | 4.77 | 849.31 |
| 29 | MCOLN2 | 0.0048333 | mucolipin TRP cation channel 2 | GTEx | DepMap | Descartes | 2.36 | 200.16 |
| 30 | FAM49A | 0.0045963 | NA | GTEx | DepMap | Descartes | 2.86 | NA |
| 31 | EHF | 0.0045781 | ETS homologous factor | GTEx | DepMap | Descartes | 0.40 | 29.14 |
| 32 | REL | 0.0045148 | REL proto-oncogene, NF-kB subunit | GTEx | DepMap | Descartes | 16.14 | 381.14 |
| 33 | SYNGR2 | 0.0044516 | synaptogyrin 2 | GTEx | DepMap | Descartes | 2.03 | 171.27 |
| 34 | TSPAN33 | 0.0044153 | tetraspanin 33 | GTEx | DepMap | Descartes | 1.16 | 92.14 |
| 35 | IL1R2 | 0.0043837 | interleukin 1 receptor type 2 | GTEx | DepMap | Descartes | 1.05 | 184.06 |
| 36 | NFKB1 | 0.0043713 | nuclear factor kappa B subunit 1 | GTEx | DepMap | Descartes | 13.99 | 913.36 |
| 37 | SAT1 | 0.0043208 | spermidine/spermine N1-acetyltransferase 1 | GTEx | DepMap | Descartes | 32.39 | 7411.72 |
| 38 | EBI3 | 0.0042778 | Epstein-Barr virus induced 3 | GTEx | DepMap | Descartes | 1.11 | 215.29 |
| 39 | CD207 | 0.0042092 | CD207 molecule | GTEx | DepMap | Descartes | 0.27 | 32.62 |
| 40 | GNA12 | 0.0041987 | G protein subunit alpha 12 | GTEx | DepMap | Descartes | 1.69 | 86.47 |
| 41 | CIITA | 0.0041705 | class II major histocompatibility complex transactivator | GTEx | DepMap | Descartes | 0.97 | 13.85 |
| 42 | FTH1 | 0.0041222 | ferritin heavy chain 1 | GTEx | DepMap | Descartes | 87.67 | 17723.15 |
| 43 | CD40 | 0.0040820 | CD40 molecule | GTEx | DepMap | Descartes | 1.55 | 234.66 |
| 44 | CXCL16 | 0.0040753 | C-X-C motif chemokine ligand 16 | GTEx | DepMap | Descartes | 2.35 | 276.17 |
| 45 | TNFAIP2 | 0.0040163 | TNF alpha induced protein 2 | GTEx | DepMap | Descartes | 11.79 | 675.88 |
| 46 | PNRC1 | 0.0039896 | proline rich nuclear receptor coactivator 1 | GTEx | DepMap | Descartes | 14.53 | 1709.06 |
| 47 | EDEM1 | 0.0039725 | ER degradation enhancing alpha-mannosidase like protein 1 | GTEx | DepMap | Descartes | 2.36 | 98.40 |
| 48 | PLD4 | 0.0039180 | phospholipase D family member 4 | GTEx | DepMap | Descartes | 1.13 | 135.01 |
| 49 | TTYH2 | 0.0037877 | tweety family member 2 | GTEx | DepMap | Descartes | 0.95 | 64.38 |
| 50 | BID | 0.0037863 | BH3 interacting domain death agonist | GTEx | DepMap | Descartes | 1.89 | 194.32 |
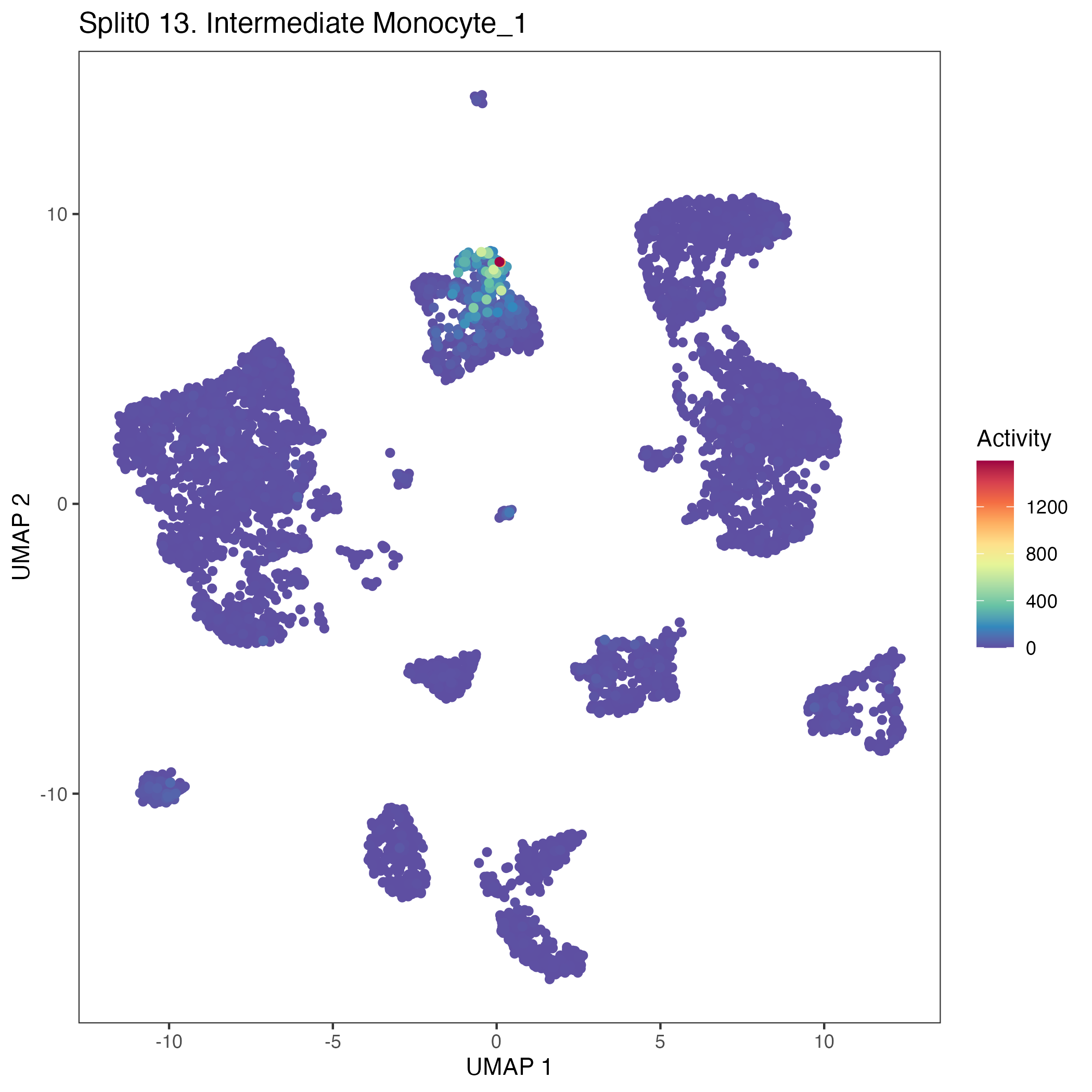
UMAP plots showing activity of gene expression program identified in community:13. Intermediate Monocyte
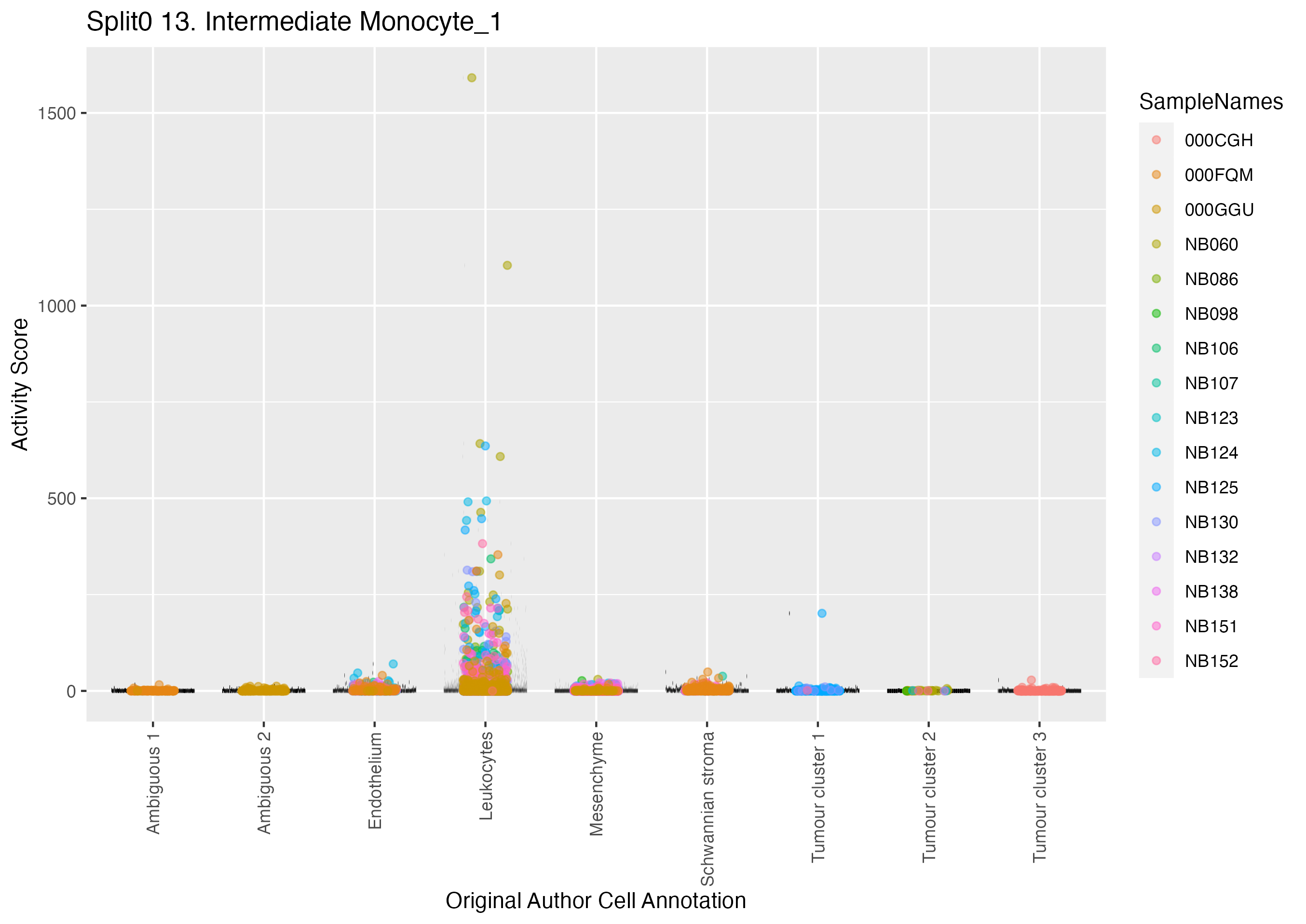
Boxlot showing activity of gene expression program identified in GEP 13. Intermediate Monocyte:
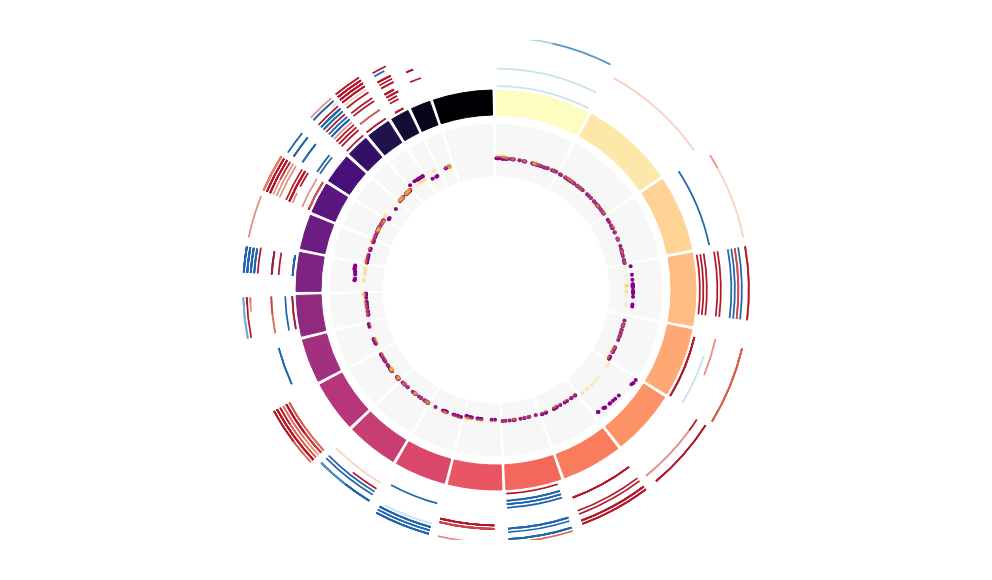
CNV Data procured from inferCNV.
Outer tracks are putative CNV regions (gains = red, losses = blue) for each patient
Inner track is expression data representing:
The top cells expressing this GEP (purple)
Random cells (n =50) from the reference set used in inferCNV (orange)
Gene set Enrichments for this program, caculated from top 50 genes
mSigDB Cell Types Gene Set:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| DESCARTES_FETAL_STOMACH_MYELOID_CELLS | 2.87e-13 | 47.27 | 20.21 | 1.92e-11 | 1.92e-10 | 10CD74, IDO1, CD83, CST3, CPVL, SPI1, WDFY4, NR4A3, CIITA, PLD4 |
76 |
| DESCARTES_FETAL_THYMUS_ANTIGEN_PRESENTING_CELLS | 2.03e-18 | 37.68 | 19.01 | 6.82e-16 | 1.36e-15 | 16CD74, IDO1, DAPP1, CD83, GPR157, CPVL, KYNU, SPI1, WDFY4, BIRC3, SLAMF7, MIR155HG, MCOLN2, TSPAN33, EBI3, PLD4 |
172 |
| DESCARTES_FETAL_INTESTINE_MYELOID_CELLS | 1.73e-19 | 33.64 | 17.49 | 1.16e-16 | 1.16e-16 | 18CD74, IDO1, DAPP1, CD83, CST3, CPVL, GPR183, KYNU, SPI1, WDFY4, SLAMF7, MCOLN2, IL1R2, EBI3, CD207, CIITA, PLD4, BID |
227 |
| AIZARANI_LIVER_C25_KUPFFER_CELLS_4 | 8.84e-17 | 33.68 | 16.74 | 1.98e-14 | 5.93e-14 | 15CD83, SERPINB9, CST3, CPVL, GPR183, OGFRL1, CMTM6, NR4A3, REL, IL1R2, SAT1, CIITA, FTH1, CXCL16, PLD4 |
174 |
| DURANTE_ADULT_OLFACTORY_NEUROEPITHELIUM_B_CELLS | 2.39e-07 | 46.13 | 13.38 | 6.42e-06 | 1.61e-04 | 5CD74, CD83, GPR183, BIRC3, REL |
35 |
| AIZARANI_LIVER_C2_KUPFFER_CELLS_1 | 1.98e-14 | 26.13 | 12.79 | 2.21e-12 | 1.33e-11 | 14CD83, CST3, CPVL, GPR183, OGFRL1, WDFY4, NR4A3, REL, IL1R2, SAT1, CIITA, FTH1, CXCL16, PLD4 |
200 |
| BUSSLINGER_ESOPHAGEAL_DENDRITIC_CELLS | 4.99e-14 | 24.29 | 11.91 | 4.79e-12 | 3.35e-11 | 14CD74, CD83, SERPINB9, CST3, CPVL, GPR183, CMTM6, SPI1, NR4A3, SYNGR2, CD207, FTH1, CXCL16, PNRC1 |
214 |
| GAO_LARGE_INTESTINE_24W_C11_PANETH_LIKE_CELL | 2.10e-15 | 20.91 | 10.80 | 2.82e-13 | 1.41e-12 | 17CD74, CD83, CPVL, OGFRL1, KYNU, SPI1, WDFY4, BIRC3, SLAMF7, NR4A3, TSPAN33, IL1R2, EBI3, CD40, CXCL16, TNFAIP2, EDEM1 |
325 |
| DURANTE_ADULT_OLFACTORY_NEUROEPITHELIUM_MACROPHAGES | 2.74e-08 | 27.47 | 10.09 | 9.95e-07 | 1.84e-05 | 7CD74, CD83, CST3, GPR183, SPI1, CXCL16, PLD4 |
81 |
| DESCARTES_FETAL_LUNG_MYELOID_CELLS | 6.06e-11 | 21.36 | 9.69 | 3.70e-09 | 4.07e-08 | 11CD74, IDO1, CD83, CPVL, KYNU, SPI1, WDFY4, IL1R2, CD207, CIITA, PLD4 |
176 |
| DESCARTES_FETAL_MUSCLE_MYELOID_CELLS | 1.66e-09 | 22.67 | 9.47 | 7.98e-08 | 1.12e-06 | 9CD74, CD83, CST3, KYNU, SPI1, MIR155HG, MCOLN2, CIITA, PLD4 |
130 |
| FAN_OVARY_CL13_MONOCYTE_MACROPHAGE | 1.87e-15 | 17.46 | 9.24 | 2.82e-13 | 1.26e-12 | 19CD74, CCR7, CD83, SERPINB9, CST3, CPVL, GPR183, KYNU, CMTM6, SPI1, G0S2, LGALS2, REL, SYNGR2, NFKB1, SAT1, FTH1, CXCL16, BID |
458 |
| DESCARTES_FETAL_PLACENTA_MYELOID_CELLS | 4.61e-09 | 20.03 | 8.39 | 2.06e-07 | 3.09e-06 | 9CD74, LAMP3, CD83, CYP2S1, CPVL, LGALS2, PADI2, CIITA, CXCL16 |
146 |
| FAN_OVARY_CL18_B_LYMPHOCYTE | 1.37e-13 | 15.91 | 8.24 | 1.02e-11 | 9.19e-11 | 17CD74, CCR7, CD83, SLC7A11, SERPINB9, CST3, GPR183, CMTM6, BIRC3, MIR155HG, REL, SYNGR2, NFKB1, SAT1, FTH1, PLD4, BID |
422 |
| DURANTE_ADULT_OLFACTORY_NEUROEPITHELIUM_MONOCYTES | 1.64e-05 | 31.00 | 7.69 | 3.43e-04 | 1.10e-02 | 4CD74, CD83, GPR183, REL |
39 |
| DESCARTES_FETAL_HEART_MYELOID_CELLS | 4.65e-08 | 18.89 | 7.50 | 1.56e-06 | 3.12e-05 | 8CD74, CYP2S1, SPI1, PADI2, IL1R2, EBI3, CIITA, PLD4 |
134 |
| TRAVAGLINI_LUNG_EREG_DENDRITIC_CELL | 1.11e-13 | 13.69 | 7.26 | 9.34e-12 | 7.47e-11 | 19CD74, CD83, SERPINB9, CST3, CPVL, GPR183, KYNU, CMTM6, SPI1, G0S2, REL, IL1R2, NFKB1, SAT1, GNA12, FTH1, CXCL16, PNRC1, EDEM1 |
579 |
| DESCARTES_FETAL_PANCREAS_MYELOID_CELLS | 1.17e-09 | 15.88 | 7.24 | 6.02e-08 | 7.82e-07 | 11IDO1, CD83, CPVL, OGFRL1, KYNU, SPI1, SLAMF7, PADI2, MCOLN2, IL1R2, PLD4 |
233 |
| DURANTE_ADULT_OLFACTORY_NEUROEPITHELIUM_DENDRITIC_CELLS | 3.45e-07 | 18.49 | 6.86 | 8.89e-06 | 2.31e-04 | 7CST3, CPVL, SPI1, G0S2, LGALS2, FTH1, TNFAIP2 |
117 |
| TRAVAGLINI_LUNG_NONCLASSICAL_MONOCYTE_CELL | 2.82e-08 | 16.04 | 6.75 | 9.95e-07 | 1.89e-05 | 9CST3, CPVL, KYNU, CMTM6, SPI1, SAT1, FTH1, CXCL16, BID |
180 |
Dowload full table
mSigDB Hallmark Gene Sets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| HALLMARK_TNFA_SIGNALING_VIA_NFKB | 2.35e-10 | 18.65 | 8.48 | 1.18e-08 | 1.18e-08 | 11CD83, GPR183, KYNU, G0S2, BIRC3, NR4A3, REL, NFKB1, SAT1, TNFAIP2, PNRC1 |
200 |
| HALLMARK_INTERFERON_GAMMA_RESPONSE | 1.19e-05 | 10.54 | 3.95 | 2.97e-04 | 5.94e-04 | 7CD74, IDO1, SLAMF7, NFKB1, CIITA, CD40, TNFAIP2 |
200 |
| HALLMARK_INFLAMMATORY_RESPONSE | 1.24e-04 | 8.78 | 3.02 | 2.06e-03 | 6.18e-03 | 6LAMP3, CCR7, GPR183, NFKB1, EBI3, CD40 |
200 |
| HALLMARK_APOPTOSIS | 2.53e-02 | 5.05 | 0.99 | 2.32e-01 | 1.00e+00 | 3BIRC3, SAT1, BID |
161 |
| HALLMARK_CHOLESTEROL_HOMEOSTASIS | 3.47e-02 | 7.23 | 0.84 | 2.32e-01 | 1.00e+00 | 2CXCL16, PNRC1 |
74 |
| HALLMARK_IL2_STAT5_SIGNALING | 4.30e-02 | 4.07 | 0.80 | 2.32e-01 | 1.00e+00 | 3CD83, SYNGR2, IL1R2 |
199 |
| HALLMARK_MTORC1_SIGNALING | 4.35e-02 | 4.05 | 0.80 | 2.32e-01 | 1.00e+00 | 3DAPP1, SLC7A11, EDEM1 |
200 |
| HALLMARK_XENOBIOTIC_METABOLISM | 4.35e-02 | 4.05 | 0.80 | 2.32e-01 | 1.00e+00 | 3CYP2S1, SLC6A12, KYNU |
200 |
| HALLMARK_ALLOGRAFT_REJECTION | 4.35e-02 | 4.05 | 0.80 | 2.32e-01 | 1.00e+00 | 3CD74, SPI1, CD40 |
200 |
| HALLMARK_IL6_JAK_STAT3_SIGNALING | 4.64e-02 | 6.13 | 0.71 | 2.32e-01 | 1.00e+00 | 2IL1R2, EBI3 |
87 |
| HALLMARK_INTERFERON_ALPHA_RESPONSE | 5.62e-02 | 5.48 | 0.64 | 2.56e-01 | 1.00e+00 | 2CD74, LAMP3 |
97 |
| HALLMARK_UV_RESPONSE_UP | 1.28e-01 | 3.34 | 0.39 | 5.34e-01 | 1.00e+00 | 2SLC6A12, BID |
158 |
| HALLMARK_P53_PATHWAY | 1.85e-01 | 2.63 | 0.31 | 6.60e-01 | 1.00e+00 | 2SLC7A11, SAT1 |
200 |
| HALLMARK_KRAS_SIGNALING_UP | 1.85e-01 | 2.63 | 0.31 | 6.60e-01 | 1.00e+00 | 2G0S2, BIRC3 |
200 |
| HALLMARK_ANDROGEN_RESPONSE | 3.28e-01 | 2.58 | 0.06 | 1.00e+00 | 1.00e+00 | 1SAT1 |
100 |
| HALLMARK_PI3K_AKT_MTOR_SIGNALING | 3.41e-01 | 2.45 | 0.06 | 1.00e+00 | 1.00e+00 | 1DAPP1 |
105 |
| HALLMARK_UNFOLDED_PROTEIN_RESPONSE | 3.62e-01 | 2.28 | 0.06 | 1.00e+00 | 1.00e+00 | 1EDEM1 |
113 |
| HALLMARK_UV_RESPONSE_DN | 4.35e-01 | 1.78 | 0.04 | 1.00e+00 | 1.00e+00 | 1NFKB1 |
144 |
| HALLMARK_FATTY_ACID_METABOLISM | 4.66e-01 | 1.62 | 0.04 | 1.00e+00 | 1.00e+00 | 1G0S2 |
158 |
| HALLMARK_HYPOXIA | 5.47e-01 | 1.28 | 0.03 | 1.00e+00 | 1.00e+00 | 1PNRC1 |
200 |
Dowload full table
KEGG Pathways:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| KEGG_APOPTOSIS | 4.89e-03 | 9.50 | 1.85 | 6.79e-01 | 9.10e-01 | 3BIRC3, NFKB1, BID |
87 |
| KEGG_PRIMARY_IMMUNODEFICIENCY | 8.47e-03 | 15.77 | 1.78 | 6.79e-01 | 1.00e+00 | 2CIITA, CD40 |
35 |
| KEGG_TRYPTOPHAN_METABOLISM | 1.10e-02 | 13.70 | 1.56 | 6.79e-01 | 1.00e+00 | 2IDO1, KYNU |
40 |
| KEGG_ACUTE_MYELOID_LEUKEMIA | 2.14e-02 | 9.46 | 1.09 | 7.04e-01 | 1.00e+00 | 2SPI1, NFKB1 |
57 |
| KEGG_CYTOKINE_CYTOKINE_RECEPTOR_INTERACTION | 1.96e-02 | 4.16 | 1.08 | 7.04e-01 | 1.00e+00 | 4CCR7, IL1R2, CD40, CXCL16 |
265 |
| KEGG_NOD_LIKE_RECEPTOR_SIGNALING_PATHWAY | 2.50e-02 | 8.68 | 1.00 | 7.04e-01 | 1.00e+00 | 2BIRC3, NFKB1 |
62 |
| KEGG_VIRAL_MYOCARDITIS | 3.13e-02 | 7.66 | 0.88 | 7.04e-01 | 1.00e+00 | 2CD40, BID |
70 |
| KEGG_PATHWAYS_IN_CANCER | 3.70e-02 | 3.39 | 0.88 | 7.04e-01 | 1.00e+00 | 4SPI1, BIRC3, NFKB1, BID |
325 |
| KEGG_CHEMOKINE_SIGNALING_PATHWAY | 3.79e-02 | 4.29 | 0.85 | 7.04e-01 | 1.00e+00 | 3CCR7, NFKB1, CXCL16 |
189 |
| KEGG_B_CELL_RECEPTOR_SIGNALING_PATHWAY | 3.55e-02 | 7.13 | 0.82 | 7.04e-01 | 1.00e+00 | 2DAPP1, NFKB1 |
75 |
| KEGG_SMALL_CELL_LUNG_CANCER | 4.36e-02 | 6.35 | 0.74 | 7.34e-01 | 1.00e+00 | 2BIRC3, NFKB1 |
84 |
| KEGG_ANTIGEN_PROCESSING_AND_PRESENTATION | 4.73e-02 | 6.05 | 0.70 | 7.34e-01 | 1.00e+00 | 2CD74, CIITA |
88 |
| KEGG_TOLL_LIKE_RECEPTOR_SIGNALING_PATHWAY | 6.14e-02 | 5.21 | 0.60 | 8.79e-01 | 1.00e+00 | 2NFKB1, CD40 |
102 |
| KEGG_MAPK_SIGNALING_PATHWAY | 8.58e-02 | 3.02 | 0.60 | 1.00e+00 | 1.00e+00 | 3IL1R2, NFKB1, GNA12 |
267 |
| KEGG_ASTHMA | 1.13e-01 | 8.79 | 0.21 | 1.00e+00 | 1.00e+00 | 1CD40 |
30 |
| KEGG_ALLOGRAFT_REJECTION | 1.37e-01 | 7.08 | 0.17 | 1.00e+00 | 1.00e+00 | 1CD40 |
37 |
| KEGG_PORPHYRIN_AND_CHLOROPHYLL_METABOLISM | 1.51e-01 | 6.37 | 0.15 | 1.00e+00 | 1.00e+00 | 1FTH1 |
41 |
| KEGG_INTESTINAL_IMMUNE_NETWORK_FOR_IGA_PRODUCTION | 1.74e-01 | 5.43 | 0.13 | 1.00e+00 | 1.00e+00 | 1CD40 |
48 |
| KEGG_AUTOIMMUNE_THYROID_DISEASE | 1.87e-01 | 5.00 | 0.12 | 1.00e+00 | 1.00e+00 | 1CD40 |
52 |
| KEGG_AMYOTROPHIC_LATERAL_SCLEROSIS_ALS | 1.90e-01 | 4.91 | 0.12 | 1.00e+00 | 1.00e+00 | 1BID |
53 |
Dowload full table
CHR Positional Gene Sets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| chr6p23 | 5.81e-02 | 18.20 | 0.42 | 1.00e+00 | 1.00e+00 | 1CD83 |
15 |
| chr1q32 | 2.78e-01 | 1.97 | 0.23 | 1.00e+00 | 1.00e+00 | 2LAD1, G0S2 |
266 |
| chr17q25 | 3.22e-01 | 1.77 | 0.21 | 1.00e+00 | 1.00e+00 | 2SYNGR2, TTYH2 |
297 |
| chr3p26 | 1.61e-01 | 5.93 | 0.14 | 1.00e+00 | 1.00e+00 | 1EDEM1 |
44 |
| chr6q15 | 1.74e-01 | 5.43 | 0.13 | 1.00e+00 | 1.00e+00 | 1PNRC1 |
48 |
| chr4q24 | 2.00e-01 | 4.64 | 0.11 | 1.00e+00 | 1.00e+00 | 1NFKB1 |
56 |
| chr6q13 | 2.00e-01 | 4.64 | 0.11 | 1.00e+00 | 1.00e+00 | 1OGFRL1 |
56 |
| chr14q32 | 1.00e+00 | 0.96 | 0.11 | 1.00e+00 | 1.00e+00 | 2TNFAIP2, PLD4 |
546 |
| chr1p36 | 1.00e+00 | 0.80 | 0.09 | 1.00e+00 | 1.00e+00 | 2GPR157, PADI2 |
656 |
| chr2q22 | 2.37e-01 | 3.81 | 0.09 | 1.00e+00 | 1.00e+00 | 1KYNU |
68 |
| chr6p25 | 2.78e-01 | 3.15 | 0.08 | 1.00e+00 | 1.00e+00 | 1SERPINB9 |
82 |
| chr7q32 | 3.01e-01 | 2.87 | 0.07 | 1.00e+00 | 1.00e+00 | 1TSPAN33 |
90 |
| chr13q32 | 3.15e-01 | 2.71 | 0.07 | 1.00e+00 | 1.00e+00 | 1GPR183 |
95 |
| chr8p11 | 3.15e-01 | 2.71 | 0.07 | 1.00e+00 | 1.00e+00 | 1IDO1 |
95 |
| chr11q22 | 3.23e-01 | 2.63 | 0.06 | 1.00e+00 | 1.00e+00 | 1BIRC3 |
98 |
| chr5q33 | 3.52e-01 | 2.36 | 0.06 | 1.00e+00 | 1.00e+00 | 1CD74 |
109 |
| chr2p16 | 3.64e-01 | 2.26 | 0.06 | 1.00e+00 | 1.00e+00 | 1REL |
114 |
| chr3q27 | 3.67e-01 | 2.24 | 0.06 | 1.00e+00 | 1.00e+00 | 1LAMP3 |
115 |
| chr21q21 | 3.77e-01 | 2.16 | 0.05 | 1.00e+00 | 1.00e+00 | 1MIR155HG |
119 |
| chr7p22 | 3.82e-01 | 2.13 | 0.05 | 1.00e+00 | 1.00e+00 | 1GNA12 |
121 |
Dowload full table
Transcription Factor Targets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| NFKAPPAB_01 | 5.50e-05 | 8.21 | 3.08 | 6.23e-02 | 6.23e-02 | 7CD83, SLC6A12, BIRC3, EHF, EBI3, CD40, CXCL16 |
255 |
| NFKAPPAB65_01 | 3.34e-04 | 7.25 | 2.50 | 1.89e-01 | 3.78e-01 | 6SLC6A12, BIRC3, EHF, REL, CD40, CXCL16 |
241 |
| GGGNNTTTCC_NFKB_Q6_01 | 1.88e-03 | 8.36 | 2.15 | 2.92e-01 | 1.00e+00 | 4CD74, SLC6A12, BIRC3, REL |
134 |
| MAML1_TARGET_GENES | 1.26e-03 | 5.57 | 1.93 | 2.92e-01 | 1.00e+00 | 6BIRC3, MIR155HG, REL, SAT1, EBI3, CIITA |
312 |
| NFKB_Q6_01 | 2.25e-03 | 5.99 | 1.84 | 2.92e-01 | 1.00e+00 | 5SLC6A12, BIRC3, EHF, EBI3, CD40 |
237 |
| ICSBP_Q6 | 2.97e-03 | 5.60 | 1.72 | 2.92e-01 | 1.00e+00 | 5IDO1, DAPP1, KYNU, NFKB1, CXCL16 |
253 |
| CREL_01 | 3.18e-03 | 5.51 | 1.69 | 2.92e-01 | 1.00e+00 | 5CD74, SLC6A12, BIRC3, EHF, REL |
257 |
| NFKB_Q6 | 3.23e-03 | 5.49 | 1.69 | 2.92e-01 | 1.00e+00 | 5CD83, SLC6A12, EHF, REL, CD40 |
258 |
| RYTTCCTG_ETS2_B | 6.26e-03 | 2.84 | 1.26 | 5.06e-01 | 1.00e+00 | 10DAPP1, GPR183, RASAL1, CMTM6, SPI1, WDFY4, NR4A3, NFKB1, FTH1, CD40 |
1112 |
| IRF_Q6 | 1.50e-02 | 4.53 | 1.17 | 7.07e-01 | 1.00e+00 | 4DAPP1, KYNU, SAT1, CIITA |
244 |
| IRF7_01 | 1.80e-02 | 4.28 | 1.11 | 7.12e-01 | 1.00e+00 | 4DAPP1, NR4A3, SAT1, CXCL16 |
258 |
| ZNF597_TARGET_GENES | 1.49e-02 | 2.74 | 1.11 | 7.07e-01 | 1.00e+00 | 8CCR7, CD83, BIRC3, MIR155HG, REL, SYNGR2, NFKB1, EBI3 |
877 |
| IRF1_Q6 | 1.91e-02 | 4.20 | 1.09 | 7.12e-01 | 1.00e+00 | 4CCR7, DAPP1, KYNU, CIITA |
263 |
| CGTSACG_PAX3_B | 2.08e-02 | 5.46 | 1.08 | 7.12e-01 | 1.00e+00 | 3SLC6A12, PNRC1, EDEM1 |
149 |
| NFKB_C | 2.03e-02 | 4.12 | 1.07 | 7.12e-01 | 1.00e+00 | 4CD74, SLC6A12, EHF, CD40 |
268 |
| CCAATNNSNNNGCG_UNKNOWN | 2.36e-02 | 8.98 | 1.03 | 7.63e-01 | 1.00e+00 | 2PNRC1, EDEM1 |
60 |
| ZNF768_TARGET_GENES | 5.47e-02 | 2.05 | 0.88 | 9.56e-01 | 1.00e+00 | 9CD83, SLC7A11, SLC6A12, CPVL, GPR183, RASAL1, MIR155HG, CIITA, FTH1 |
1346 |
| NCOA2_TARGET_GENES | 5.09e-02 | 2.65 | 0.82 | 9.56e-01 | 1.00e+00 | 5OGFRL1, SYNGR2, EBI3, CIITA, CXCL16 |
530 |
| SMTTTTGT_UNKNOWN | 7.14e-02 | 2.70 | 0.70 | 9.56e-01 | 1.00e+00 | 4CCR7, OGFRL1, NR4A3, EHF |
407 |
| IK3_01 | 6.02e-02 | 3.53 | 0.70 | 9.56e-01 | 1.00e+00 | 3EHF, CD40, EDEM1 |
229 |
Dowload full table
GO Biological Processes:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| GOBP_TRYPTOPHAN_CATABOLIC_PROCESS_TO_KYNURENINE | 1.54e-04 | 172.68 | 14.09 | 1.44e-01 | 1.00e+00 | 2IDO1, KYNU |
5 |
| GOBP_POSITIVE_REGULATION_OF_DENDRITIC_CELL_ANTIGEN_PROCESSING_AND_PRESENTATION | 3.22e-04 | 103.49 | 9.64 | 1.72e-01 | 1.00e+00 | 2CD74, CCR7 |
7 |
| GOBP_DE_NOVO_NAD_BIOSYNTHETIC_PROCESS | 4.28e-04 | 86.37 | 8.33 | 1.89e-01 | 1.00e+00 | 2IDO1, KYNU |
8 |
| GOBP_POSITIVE_REGULATION_OF_ANTIGEN_PROCESSING_AND_PRESENTATION | 5.49e-04 | 74.10 | 7.33 | 1.96e-01 | 1.00e+00 | 2CD74, CCR7 |
9 |
| GOBP_TRYPTOPHAN_CATABOLIC_PROCESS | 6.85e-04 | 64.86 | 6.55 | 2.05e-01 | 1.00e+00 | 2IDO1, KYNU |
10 |
| GOBP_POSITIVE_REGULATION_OF_CHEMOKINE_C_X_C_MOTIF_LIGAND_2_PRODUCTION | 6.85e-04 | 64.86 | 6.55 | 2.05e-01 | 1.00e+00 | 2CD74, MCOLN2 |
10 |
| GOBP_REGULATION_OF_DENDRITIC_CELL_ANTIGEN_PROCESSING_AND_PRESENTATION | 8.35e-04 | 57.69 | 5.91 | 2.08e-01 | 1.00e+00 | 2CD74, CCR7 |
11 |
| GOBP_KYNURENINE_METABOLIC_PROCESS | 8.35e-04 | 57.69 | 5.91 | 2.08e-01 | 1.00e+00 | 2IDO1, KYNU |
11 |
| GOBP_TRYPTOPHAN_METABOLIC_PROCESS | 9.99e-04 | 51.89 | 5.39 | 2.20e-01 | 1.00e+00 | 2IDO1, KYNU |
12 |
| GOBP_NEGATIVE_T_CELL_SELECTION | 9.99e-04 | 51.89 | 5.39 | 2.20e-01 | 1.00e+00 | 2CD74, CCR7 |
12 |
| GOBP_AMINE_CATABOLIC_PROCESS | 2.70e-04 | 27.48 | 5.18 | 1.62e-01 | 1.00e+00 | 3IDO1, KYNU, SAT1 |
32 |
| GOBP_INTERLEUKIN_12_PRODUCTION | 9.06e-05 | 19.39 | 4.91 | 9.68e-02 | 6.78e-01 | 4IDO1, CCR7, NFKB1, CD40 |
60 |
| GOBP_DENDRITIC_CELL_ANTIGEN_PROCESSING_AND_PRESENTATION | 1.58e-03 | 39.95 | 4.27 | 3.02e-01 | 1.00e+00 | 2CD74, CCR7 |
15 |
| GOBP_NEGATIVE_REGULATION_OF_COLLAGEN_METABOLIC_PROCESS | 1.58e-03 | 39.95 | 4.27 | 3.02e-01 | 1.00e+00 | 2CST3, CIITA |
15 |
| GOBP_POSITIVE_REGULATION_OF_B_CELL_PROLIFERATION | 5.25e-04 | 21.53 | 4.11 | 1.96e-01 | 1.00e+00 | 3CD74, GPR183, CD40 |
40 |
| GOBP_POSITIVE_REGULATION_OF_INTERLEUKIN_12_PRODUCTION | 5.25e-04 | 21.53 | 4.11 | 1.96e-01 | 1.00e+00 | 3IDO1, CCR7, CD40 |
40 |
| GOBP_POSITIVE_REGULATION_OF_TYPE_2_IMMUNE_RESPONSE | 1.80e-03 | 37.11 | 3.99 | 3.07e-01 | 1.00e+00 | 2CD74, IDO1 |
16 |
| GOBP_POSITIVE_REGULATION_OF_MYELOID_LEUKOCYTE_CYTOKINE_PRODUCTION_INVOLVED_IN_IMMUNE_RESPONSE | 1.80e-03 | 37.11 | 3.99 | 3.07e-01 | 1.00e+00 | 2CD74, NR4A3 |
16 |
| GOBP_NEGATIVE_REGULATION_OF_OXIDATIVE_STRESS_INDUCED_NEURON_DEATH | 1.80e-03 | 37.11 | 3.99 | 3.07e-01 | 1.00e+00 | 2SLC7A11, NR4A3 |
16 |
| GOBP_INDOLALKYLAMINE_METABOLIC_PROCESS | 2.03e-03 | 34.68 | 3.75 | 3.12e-01 | 1.00e+00 | 2IDO1, KYNU |
17 |
Dowload full table
Immunological Gene Sets:
| P-value | OR | Lower 95% CI | FDR | FWER | Genes Found | Gene Set Size | |
|---|---|---|---|---|---|---|---|
| GSE42724_MEMORY_VS_B1_BCELL_DN | 5.62e-16 | 29.42 | 14.66 | 2.74e-12 | 2.74e-12 | 15IDO1, CCR7, SLC7A11, LAD1, KYNU, MCOLN2, NFKB1, SAT1, EBI3, FTH1, CD40, TNFAIP2, PNRC1, EDEM1, TTYH2 |
197 |
| GSE2706_UNSTIM_VS_8H_R848_DC_DN | 1.61e-14 | 26.55 | 13.00 | 3.91e-11 | 7.83e-11 | 14CCR7, GPR157, G0S2, BIRC3, SLAMF7, NR4A3, MIR155HG, MCOLN2, REL, NFKB1, SAT1, EBI3, TNFAIP2, EDEM1 |
197 |
| GSE2706_UNSTIM_VS_2H_LPS_AND_R848_DC_DN | 2.80e-13 | 24.68 | 11.82 | 4.55e-10 | 1.37e-09 | 13LAMP3, CCR7, DAPP1, GPR183, BIRC3, SLAMF7, NR4A3, MIR155HG, REL, NFKB1, TNFAIP2, TTYH2, BID |
191 |
| GSE19198_1H_VS_24H_IL21_TREATED_TCELL_DN | 5.03e-13 | 23.47 | 11.26 | 6.12e-10 | 2.45e-09 | 13CCR7, CD83, SLC7A11, KYNU, LGALS2, BIRC3, SLAMF7, NR4A3, EHF, TSPAN33, CD40, TTYH2, BID |
200 |
| GSE41087_WT_VS_FOXP3_MUT_ANTI_CD3_CD28_STIM_CD4_TCELL_UP | 8.57e-12 | 21.58 | 10.09 | 8.00e-09 | 4.17e-08 | 12IDO1, LAMP3, CCR7, CD83, SLC7A11, GPR183, KYNU, BIRC3, SLAMF7, NFKB1, PNRC1, BID |
195 |
| GSE19198_6H_VS_24H_IL21_TREATED_TCELL_UP | 1.08e-11 | 21.10 | 9.88 | 8.00e-09 | 5.29e-08 | 12CCR7, CD83, SLC7A11, G0S2, LGALS2, BIRC3, SLAMF7, NR4A3, EHF, TSPAN33, CD40, BID |
199 |
| GSE22886_CTRL_VS_LPS_24H_DC_DN | 1.15e-11 | 20.99 | 9.83 | 8.00e-09 | 5.60e-08 | 12IDO1, LAMP3, CCR7, CD83, KYNU, G0S2, BIRC3, SLAMF7, NR4A3, REL, NFKB1, EBI3 |
200 |
| GSE2706_UNSTIM_VS_2H_R848_DC_DN | 1.62e-10 | 19.37 | 8.80 | 9.84e-08 | 7.87e-07 | 11SERPINB9, BIRC3, SLAMF7, NR4A3, MIR155HG, MCOLN2, REL, NFKB1, TNFAIP2, TTYH2, BID |
193 |
| GSE26343_UNSTIM_VS_LPS_STIM_NFAT5_KO_MACROPHAGE_DN | 2.35e-10 | 18.65 | 8.48 | 1.27e-07 | 1.15e-06 | 11DAPP1, CD83, SERPINB9, SLAMF7, NR4A3, MCOLN2, TSPAN33, CD40, CXCL16, TNFAIP2, PNRC1 |
200 |
| GSE2706_UNSTIM_VS_2H_LPS_DC_DN | 3.21e-09 | 16.98 | 7.45 | 1.23e-06 | 1.56e-05 | 10LAMP3, GPR183, BIRC3, NR4A3, MIR155HG, REL, NFKB1, TNFAIP2, TTYH2, BID |
194 |
| GSE9988_LPS_VS_VEHICLE_TREATED_MONOCYTE_UP | 3.54e-09 | 16.80 | 7.38 | 1.23e-06 | 1.72e-05 | 10LAMP3, CCR7, CD83, G0S2, BIRC3, SLAMF7, MIR155HG, REL, NFKB1, TNFAIP2 |
196 |
| GSE9988_LOW_LPS_VS_VEHICLE_TREATED_MONOCYTE_UP | 3.54e-09 | 16.80 | 7.38 | 1.23e-06 | 1.72e-05 | 10LAMP3, CCR7, CD83, G0S2, BIRC3, SLAMF7, MIR155HG, REL, NFKB1, TNFAIP2 |
196 |
| GSE14000_UNSTIM_VS_16H_LPS_DC_TRANSLATED_RNA_DN | 3.72e-09 | 16.71 | 7.34 | 1.23e-06 | 1.81e-05 | 10IDO1, GPR157, BIRC3, SLAMF7, NR4A3, MIR155HG, EBI3, TNFAIP2, PNRC1, TTYH2 |
197 |
| GSE46242_TH1_VS_ANERGIC_TH1_CD4_TCELL_WITH_EGR2_DELETED_UP | 4.09e-09 | 16.53 | 7.26 | 1.23e-06 | 1.99e-05 | 10IDO1, LAMP3, CCR7, CD83, GPR157, G0S2, BIRC3, SAT1, CD40, TNFAIP2 |
199 |
| GSE32986_CURDLAN_HIGHDOSE_VS_GMCSF_AND_CURDLAN_HIGHDOSE_STIM_DC_DN | 4.29e-09 | 16.45 | 7.22 | 1.23e-06 | 2.09e-05 | 10CCR7, CD83, SERPINB9, BIRC3, NR4A3, REL, NFKB1, CIITA, CD40, TNFAIP2 |
200 |
| GSE26343_WT_VS_NFAT5_KO_MACROPHAGE_DN | 4.29e-09 | 16.45 | 7.22 | 1.23e-06 | 2.09e-05 | 10DAPP1, LAD1, GPR183, RASAL1, SLAMF7, NR4A3, EBI3, TNFAIP2, PNRC1, EDEM1 |
200 |
| GSE35825_UNTREATED_VS_IFNG_STIM_MACROPHAGE_UP | 4.29e-09 | 16.45 | 7.22 | 1.23e-06 | 2.09e-05 | 10CD83, SLC7A11, LAD1, GPR183, BIRC3, SLAMF7, NR4A3, MCOLN2, EBI3, PNRC1 |
200 |
| GSE360_CTRL_VS_T_GONDII_DC_DN | 6.34e-08 | 14.52 | 6.11 | 1.46e-05 | 3.09e-04 | 9LAMP3, CCR7, CD83, NR4A3, NFKB1, SAT1, GNA12, TNFAIP2, PNRC1 |
198 |
| GSE360_T_GONDII_VS_B_MALAYI_HIGH_DOSE_DC_UP | 6.34e-08 | 14.52 | 6.11 | 1.46e-05 | 3.09e-04 | 9LAMP3, CCR7, CD83, NR4A3, REL, NFKB1, SAT1, TNFAIP2, PNRC1 |
198 |
| GSE9988_ANTI_TREM1_VS_LPS_MONOCYTE_DN | 6.34e-08 | 14.52 | 6.11 | 1.46e-05 | 3.09e-04 | 9CD83, G0S2, BIRC3, MCOLN2, NFKB1, EBI3, CD40, TNFAIP2, PNRC1 |
198 |
Top Ranked Transcription Factors for this Gene Expression Program:
| Gene Symbol | TF Rank | DNA Binding Domain | Motif Status | IUPAC PWM | GTEx | DepMap | Decartes |
|---|---|---|---|---|---|---|---|
| SPI1 | 20 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| NR4A3 | 26 | Yes | Known motif | Monomer or homomultimer | In vivo/Misc source | Only known motifs are from Transfac or HocoMoco - origin is uncertain | None |
| EHF | 31 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| REL | 32 | Yes | Known motif | Monomer or homomultimer | In vivo/Misc source | None | None |
| NFKB1 | 36 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| CIITA | 41 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | Chromatin modifier |
| CD40 | 43 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | Cell surface receptor of TNF-family |
| ETV3 | 53 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| TRAF1 | 55 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | TRAF1 is an NF-kappaB interactor (PMID: 10692572), and is unlikely to have DNA-binding activity |
| CUL1 | 69 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | Component of the E3-ubiquitin ligase complex (PMID: 11961546) |
| ETV6 | 72 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| SPIB | 73 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| HCK | 75 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | Inhibits TP73-mediated transcription activation (PMID: 17535448) |
| E2F7 | 78 | Yes | Known motif | Monomer or homomultimer | High-throughput in vitro | None | None |
| MALT1 | 92 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | Protein that operates upstream of NFKB in a signaling cascade (PMID: 18223652) |
| NFKBID | 95 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | None |
| ADAM8 | 98 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | None |
| ID2 | 99 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | ID bHLH proteins lack the basic region and should not be able to bind DNA. The HT-SELEX motif for ID4 is likely by a co-precipitated protein or it is a contamination |
| CFLAR | 103 | No | Unlikely to be sequence specific TF | Not a DNA binding protein | No motif | None | None |
| JARID2 | 105 | No | Unlikely to be sequence specific TF | Low specificity DNA-binding protein | No motif | None | Only ARID3 and ARID5 family members have sequence specificity (PMID: 15640446). |
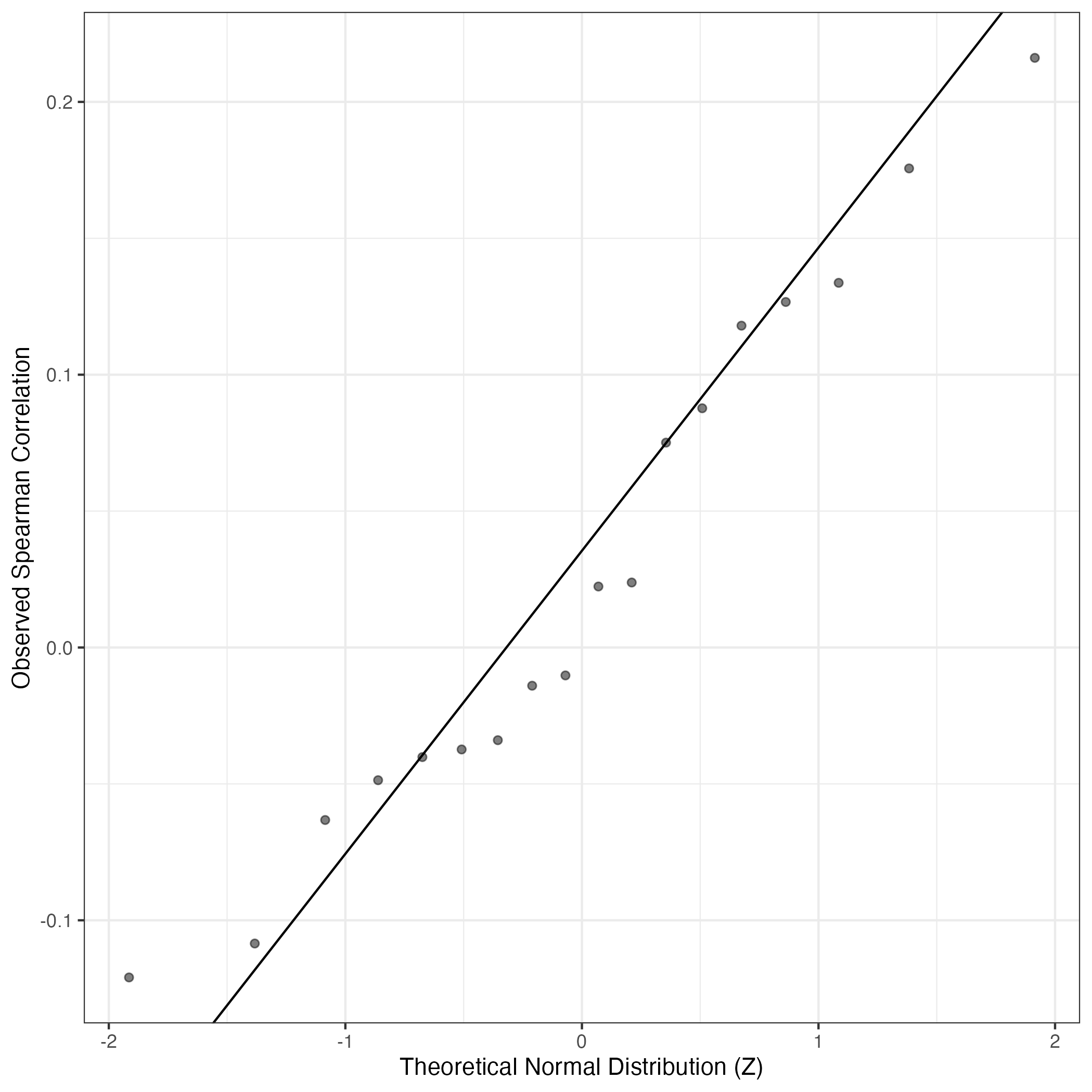
QQ Plot showing correlations with other GEPs in this dataset, calculated by Spearman correlation:
Interactive QQ-plot of gene loadings:
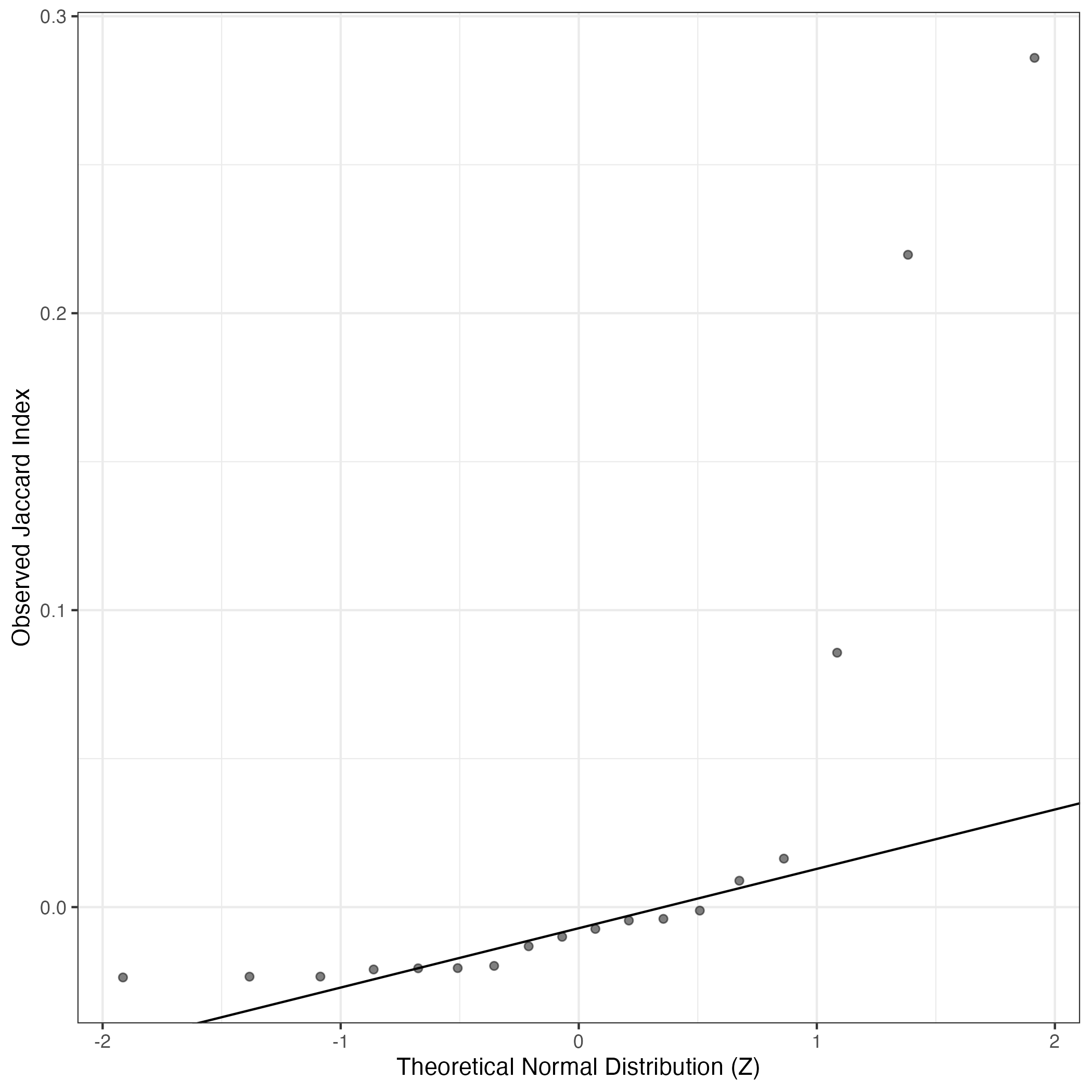
A similar QQ-plot as above, but only for instances where the H value is e.g. > 25, i.e. we are confident that the expression program is active above noise. Agreemenet between these binary vectors is tested using the Jaccard Index, with the P-values calculated by an exact test:
Interactive QQ-plot:
Singler cell type annotations for the top 50 cells on this program.
| Cell ID | Singler label | Singler Delta | Activity Score | Top Singler Raw Scores |
|---|---|---|---|---|
| TM36-O4 | Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys | 0.17 | 1591.36 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.45, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.45, Monocyte:F._tularensis_novicida: 0.45, DC:monocyte-derived:antiCD40/VAF347: 0.44, DC:monocyte-derived:Schuler_treatment: 0.44, DC:monocyte-derived:CD40L: 0.43, Monocyte:S._typhimurium_flagellin: 0.43, DC:monocyte-derived:LPS: 0.43, DC:monocyte-derived:mature: 0.43, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.43 |
| TM38-M8 | DC:monocyte-derived:mature | 0.16 | 1104.51 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.47, DC:monocyte-derived:antiCD40/VAF347: 0.47, Monocyte:S._typhimurium_flagellin: 0.47, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.47, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.46, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.46, DC:monocyte-derived:CD40L: 0.46, DC:monocyte-derived:LPS: 0.46, DC:monocyte-derived:mature: 0.45, Monocyte:F._tularensis_novicida: 0.45 |
| TM37-K7 | DC:monocyte-derived:Schuler_treatment | 0.19 | 641.87 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.48, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.47, DC:monocyte-derived:antiCD40/VAF347: 0.47, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.46, DC:monocyte-derived:CD40L: 0.46, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.46, Monocyte:S._typhimurium_flagellin: 0.45, DC:monocyte-derived:LPS: 0.45, DC:monocyte-derived:Galectin-1: 0.45, Macrophage:monocyte-derived:S._aureus: 0.44 |
| WK020-H14 | DC:monocyte-derived:mature | 0.09 | 635.76 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.47, Macrophage:monocyte-derived:M-CSF/IFNg: 0.46, DC:monocyte-derived:Poly(IC): 0.46, Monocyte:CD16+: 0.45, Monocyte:leukotriene_D4: 0.45, Monocyte:anti-FcgRIIB: 0.45, DC:monocyte-derived:mature: 0.45, Monocyte:CD16-: 0.45, Monocyte: 0.45, Neutrophil:GM-CSF_IFNg: 0.45 |
| TM36-N15 | Macrophage:monocyte-derived:M-CSF/Pam3Cys | 0.22 | 608.35 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.47, Monocyte:S._typhimurium_flagellin: 0.47, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.47, Monocyte:F._tularensis_novicida: 0.46, DC:monocyte-derived:antiCD40/VAF347: 0.46, Macrophage:monocyte-derived:S._aureus: 0.44, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.44, DC:monocyte-derived:Galectin-1: 0.44, DC:monocyte-derived:LPS: 0.44, DC:monocyte-derived:Schuler_treatment: 0.43 |
| WK014-M14 | DC:monocyte-derived:antiCD40/VAF347 | 0.19 | 493.10 | Raw ScoresMonocyte:S._typhimurium_flagellin: 0.48, DC:monocyte-derived:antiCD40/VAF347: 0.47, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.47, DC:monocyte-derived:Schuler_treatment: 0.47, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.47, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.46, DC:monocyte-derived:LPS: 0.45, DC:monocyte-derived:CD40L: 0.45, Monocyte:F._tularensis_novicida: 0.44, Macrophage:monocyte-derived:S._aureus: 0.44 |
| WK036-I2 | DC:monocyte-derived:mature | 0.13 | 490.68 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.42, DC:monocyte-derived:mature: 0.42, DC:monocyte-derived:antiCD40/VAF347: 0.41, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.41, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.41, DC:monocyte-derived:CD40L: 0.41, DC:monocyte-derived:immature: 0.41, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.41, DC:monocyte-derived:LPS: 0.4, Monocyte:S._typhimurium_flagellin: 0.4 |
| TM37-L22 | DC:monocyte-derived:mature | 0.16 | 463.49 | Raw ScoresDC:monocyte-derived:mature: 0.44, DC:monocyte-derived:Schuler_treatment: 0.44, DC:monocyte-derived:CD40L: 0.41, DC:monocyte-derived:immature: 0.39, DC:monocyte-derived:antiCD40/VAF347: 0.39, DC:monocyte-derived:Poly(IC): 0.39, DC:monocyte-derived:LPS: 0.39, DC:monocyte-derived:Galectin-1: 0.39, Monocyte:F._tularensis_novicida: 0.39, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.39 |
| WK016-P8 | Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys | 0.20 | 447.01 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.5, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.49, DC:monocyte-derived:Schuler_treatment: 0.48, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.48, DC:monocyte-derived:LPS: 0.48, DC:monocyte-derived:antiCD40/VAF347: 0.48, Monocyte:S._typhimurium_flagellin: 0.48, DC:monocyte-derived:Poly(IC): 0.48, DC:monocyte-derived:CD40L: 0.48, Macrophage:monocyte-derived:S._aureus: 0.47 |
| WK032-J1 | Macrophage:monocyte-derived:M-CSF/Pam3Cys | 0.21 | 442.40 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.51, DC:monocyte-derived:antiCD40/VAF347: 0.5, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.49, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.49, Monocyte:S._typhimurium_flagellin: 0.49, DC:monocyte-derived:Schuler_treatment: 0.48, DC:monocyte-derived:LPS: 0.48, Macrophage:monocyte-derived:S._aureus: 0.48, DC:monocyte-derived:Galectin-1: 0.47, DC:monocyte-derived:CD40L: 0.47 |
| WK020-N23 | Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys | 0.22 | 417.51 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.51, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.5, DC:monocyte-derived:antiCD40/VAF347: 0.49, DC:monocyte-derived:LPS: 0.49, DC:monocyte-derived:Schuler_treatment: 0.49, DC:monocyte-derived:Galectin-1: 0.48, Monocyte:S._typhimurium_flagellin: 0.48, DC:monocyte-derived:CD40L: 0.48, Macrophage:monocyte-derived:S._aureus: 0.48, DC:monocyte-derived:Poly(IC): 0.48 |
| WK072-C21 | Monocyte:S._typhimurium_flagellin | 0.17 | 382.29 | Raw ScoresMonocyte:S._typhimurium_flagellin: 0.47, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.46, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.46, DC:monocyte-derived:antiCD40/VAF347: 0.45, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.45, Monocyte:F._tularensis_novicida: 0.45, DC:monocyte-derived:Schuler_treatment: 0.45, DC:monocyte-derived:CD40L: 0.44, Monocyte: 0.44, DC:monocyte-derived:LPS: 0.44 |
| KK051-E11 | DC:monocyte-derived:antiCD40/VAF347 | 0.14 | 353.56 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.43, DC:monocyte-derived:antiCD40/VAF347: 0.43, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.43, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.43, Monocyte:S._typhimurium_flagellin: 0.42, Monocyte:leukotriene_D4: 0.42, DC:monocyte-derived:Schuler_treatment: 0.42, Macrophage:monocyte-derived:M-CSF: 0.42, DC:monocyte-derived:Galectin-1: 0.42, DC:monocyte-derived:CD40L: 0.42 |
| WMK004-B18 | DC:monocyte-derived:mature | 0.15 | 342.71 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.42, DC:monocyte-derived:mature: 0.42, DC:monocyte-derived:CD40L: 0.41, DC:monocyte-derived:LPS: 0.41, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.41, DC:monocyte-derived:Poly(IC): 0.41, DC:monocyte-derived:antiCD40/VAF347: 0.41, DC:monocyte-derived:immature: 0.4, DC:monocyte-derived:Galectin-1: 0.4, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.4 |
| WK024-H8 | Monocyte:CD16+ | 0.14 | 313.50 | Raw ScoresMonocyte:leukotriene_D4: 0.48, Monocyte:CD16+: 0.47, Monocyte: 0.47, Macrophage:monocyte-derived:M-CSF/IFNg: 0.47, Monocyte:anti-FcgRIIB: 0.47, Monocyte:CD14+: 0.46, HSC_-G-CSF: 0.46, Monocyte:CD16-: 0.46, Pre-B_cell_CD34-: 0.46, Macrophage:monocyte-derived:M-CSF: 0.45 |
| KK059-F17 | DC:monocyte-derived:mature | 0.09 | 310.99 | Raw ScoresDC:monocyte-derived:mature: 0.39, DC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:CD40L: 0.37, Monocyte:F._tularensis_novicida: 0.36, DC:monocyte-derived:Poly(IC): 0.35, B_cell:Memory: 0.35, DC:monocyte-derived:Galectin-1: 0.35, DC:monocyte-derived:anti-DC-SIGN_2h: 0.35, DC:monocyte-derived:LPS: 0.35, DC:monocyte-derived:immature: 0.35 |
| WK072-A16 | DC:monocyte-derived:antiCD40/VAF347 | 0.18 | 310.87 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.43, Monocyte:S._typhimurium_flagellin: 0.43, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.43, DC:monocyte-derived:antiCD40/VAF347: 0.43, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.43, DC:monocyte-derived:Schuler_treatment: 0.42, DC:monocyte-derived:CD40L: 0.42, DC:monocyte-derived:LPS: 0.42, Macrophage:monocyte-derived:S._aureus: 0.41, Monocyte:leukotriene_D4: 0.41 |
| TM37-E3 | DC:monocyte-derived:antiCD40/VAF347 | 0.20 | 310.61 | Raw ScoresMonocyte:S._typhimurium_flagellin: 0.46, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.46, DC:monocyte-derived:antiCD40/VAF347: 0.45, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.45, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.44, Monocyte:F._tularensis_novicida: 0.44, DC:monocyte-derived:Schuler_treatment: 0.43, DC:monocyte-derived:LPS: 0.43, Macrophage:monocyte-derived:S._aureus: 0.43, DC:monocyte-derived:CD40L: 0.43 |
| WK020-M11 | DC:monocyte-derived:Poly(IC) | 0.16 | 310.08 | Raw ScoresDC:monocyte-derived:Poly(IC): 0.4, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.4, DC:monocyte-derived:LPS: 0.4, Monocyte:F._tularensis_novicida: 0.39, DC:monocyte-derived:mature: 0.38, Monocyte:S._typhimurium_flagellin: 0.38, Macrophage:monocyte-derived:S._aureus: 0.38, DC:monocyte-derived:antiCD40/VAF347: 0.38, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.38, DC:monocyte-derived:CD40L: 0.37 |
| WK021-G3 | Monocyte:CD16+ | 0.14 | 309.56 | Raw ScoresMonocyte:CD16+: 0.49, Monocyte:CD16-: 0.48, Monocyte:leukotriene_D4: 0.48, Monocyte: 0.48, Monocyte:CD14+: 0.48, Pre-B_cell_CD34-: 0.47, Monocyte:anti-FcgRIIB: 0.47, DC:monocyte-derived:anti-DC-SIGN_2h: 0.46, Neutrophil:commensal_E._coli_MG1655: 0.46, Macrophage:monocyte-derived:M-CSF/IFNg: 0.46 |
| KK057-F17 | T_cell:CD4+ | 0.08 | 300.91 | Raw ScoresMonocyte:leukotriene_D4: 0.51, DC:monocyte-derived:anti-DC-SIGN_2h: 0.51, Monocyte: 0.51, Monocyte:CD16+: 0.51, Pre-B_cell_CD34-: 0.51, Monocyte:CD16-: 0.51, NK_cell: 0.51, Monocyte:CD14+: 0.5, B_cell:Memory: 0.5, Macrophage:monocyte-derived:M-CSF/IFNg: 0.5 |
| WK019-E22 | B_cell:Memory | 0.08 | 272.37 | Raw ScoresNK_cell: 0.49, NK_cell:IL2: 0.48, T_cell:CD4+_effector_memory: 0.48, T_cell:CD4+_central_memory: 0.47, Monocyte:leukotriene_D4: 0.47, DC:monocyte-derived:anti-DC-SIGN_2h: 0.47, T_cell:gamma-delta: 0.47, Monocyte:anti-FcgRIIB: 0.47, T_cell:CD4+: 0.47, Monocyte: 0.47 |
| WK019-O1 | DC:monocyte-derived:mature | 0.12 | 260.57 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.38, Monocyte:leukotriene_D4: 0.38, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.37, Monocyte: 0.37, Monocyte:anti-FcgRIIB: 0.37, Monocyte:S._typhimurium_flagellin: 0.37, DC:monocyte-derived:CD40L: 0.37, DC:monocyte-derived:Poly(IC): 0.37, DC:monocyte-derived:anti-DC-SIGN_2h: 0.37 |
| TM38-N10 | Monocyte:S._typhimurium_flagellin | 0.15 | 255.27 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.44, Monocyte:S._typhimurium_flagellin: 0.43, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.43, DC:monocyte-derived:antiCD40/VAF347: 0.43, DC:monocyte-derived:Schuler_treatment: 0.43, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.43, DC:monocyte-derived:CD40L: 0.42, DC:monocyte-derived:mature: 0.42, Monocyte:leukotriene_D4: 0.42, Monocyte: 0.42 |
| WK038-F11 | DC:monocyte-derived:A._fumigatus_germ_tubes_6h | 0.15 | 251.66 | Raw ScoresDC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.4, DC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:antiCD40/VAF347: 0.39, Monocyte:F._tularensis_novicida: 0.39, Monocyte:S._typhimurium_flagellin: 0.39, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.39, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.39, Monocyte:leukotriene_D4: 0.39, DC:monocyte-derived:CD40L: 0.38, DC:monocyte-derived:mature: 0.38 |
| TM37-J19 | DC:monocyte-derived:antiCD40/VAF347 | 0.18 | 249.15 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.43, DC:monocyte-derived:antiCD40/VAF347: 0.42, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.42, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.42, DC:monocyte-derived:Schuler_treatment: 0.42, Monocyte:S._typhimurium_flagellin: 0.41, DC:monocyte-derived:LPS: 0.41, DC:monocyte-derived:CD40L: 0.41, DC:monocyte-derived:Galectin-1: 0.4, Monocyte:F._tularensis_novicida: 0.4 |
| WK073-I12 | DC:monocyte-derived:antiCD40/VAF347 | 0.19 | 244.56 | Raw ScoresDC:monocyte-derived:antiCD40/VAF347: 0.45, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.45, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.44, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.44, Monocyte:S._typhimurium_flagellin: 0.44, DC:monocyte-derived:LPS: 0.43, DC:monocyte-derived:CD40L: 0.43, DC:monocyte-derived:Schuler_treatment: 0.43, Macrophage:monocyte-derived:S._aureus: 0.43, DC:monocyte-derived:Poly(IC): 0.42 |
| WK016-P7 | NK_cell | 0.08 | 239.21 | Raw ScoresMonocyte:leukotriene_D4: 0.39, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.39, DC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:mature: 0.39, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.39, Monocyte: 0.38, Monocyte:S._typhimurium_flagellin: 0.38, Monocyte:anti-FcgRIIB: 0.38, DC:monocyte-derived:anti-DC-SIGN_2h: 0.38, DC:monocyte-derived:LPS: 0.38 |
| TM37-G6 | Monocyte:S._typhimurium_flagellin | 0.16 | 235.34 | Raw ScoresDC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.38, Monocyte:S._typhimurium_flagellin: 0.38, DC:monocyte-derived:Schuler_treatment: 0.38, DC:monocyte-derived:antiCD40/VAF347: 0.38, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.37, DC:monocyte-derived:CD40L: 0.37, DC:monocyte-derived:LPS: 0.37, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.37, Monocyte:F._tularensis_novicida: 0.36, DC:monocyte-derived:mature: 0.36 |
| TM38-D21 | DC:monocyte-derived:antiCD40/VAF347 | 0.20 | 231.13 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.46, DC:monocyte-derived:antiCD40/VAF347: 0.46, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.45, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.45, Monocyte:S._typhimurium_flagellin: 0.45, DC:monocyte-derived:LPS: 0.44, Monocyte:F._tularensis_novicida: 0.43, DC:monocyte-derived:Galectin-1: 0.43, Macrophage:monocyte-derived:S._aureus: 0.43, DC:monocyte-derived:Schuler_treatment: 0.43 |
| WK024-C18 | DC:monocyte-derived:AM580 | 0.10 | 229.36 | Raw ScoresDC:monocyte-derived:AM580: 0.38, Macrophage:monocyte-derived:M-CSF/IFNg: 0.38, DC:monocyte-derived:anti-DC-SIGN_2h: 0.38, DC:monocyte-derived: 0.38, Monocyte: 0.38, Monocyte:leukotriene_D4: 0.38, DC:monocyte-derived:AEC-conditioned: 0.38, Monocyte:CD16+: 0.37, Monocyte:CD14+: 0.37, DC:monocyte-derived:mature: 0.37 |
| KK058-D3 | Monocyte:CD16+ | 0.13 | 227.20 | Raw ScoresMonocyte:leukotriene_D4: 0.52, Monocyte: 0.51, Monocyte:CD16+: 0.5, Monocyte:CD16-: 0.5, Monocyte:anti-FcgRIIB: 0.5, Monocyte:CD14+: 0.49, Macrophage:monocyte-derived:M-CSF: 0.49, Pre-B_cell_CD34-: 0.49, DC:monocyte-derived:anti-DC-SIGN_2h: 0.49, Macrophage:monocyte-derived:M-CSF/IFNg: 0.49 |
| WMK004-N13 | Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys | 0.21 | 217.09 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.51, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.5, DC:monocyte-derived:antiCD40/VAF347: 0.5, Monocyte:S._typhimurium_flagellin: 0.49, DC:monocyte-derived:LPS: 0.49, DC:monocyte-derived:Schuler_treatment: 0.49, DC:monocyte-derived:Galectin-1: 0.49, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.49, Macrophage:monocyte-derived:S._aureus: 0.48, DC:monocyte-derived:CD40L: 0.48 |
| TM38-I20 | DC:monocyte-derived:mature | 0.13 | 216.80 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.36, DC:monocyte-derived:antiCD40/VAF347: 0.36, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.35, DC:monocyte-derived:CD40L: 0.35, Monocyte:S._typhimurium_flagellin: 0.35, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.35, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.35, DC:monocyte-derived:LPS: 0.35, DC:monocyte-derived:mature: 0.35, DC:monocyte-derived:Poly(IC): 0.34 |
| WK049-E15 | Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys | 0.13 | 215.74 | Raw ScoresMonocyte:S._typhimurium_flagellin: 0.35, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.35, DC:monocyte-derived:antiCD40/VAF347: 0.34, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.34, DC:monocyte-derived:Schuler_treatment: 0.34, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.34, DC:monocyte-derived:CD40L: 0.33, DC:monocyte-derived:LPS: 0.33, Monocyte:F._tularensis_novicida: 0.33, DC:monocyte-derived:mature: 0.33 |
| WK100-J7 | Monocyte:CD16- | 0.15 | 214.99 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.41, Monocyte:S._typhimurium_flagellin: 0.41, Monocyte:CD16-: 0.41, Monocyte: 0.41, Monocyte:anti-FcgRIIB: 0.41, Monocyte:leukotriene_D4: 0.41, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.41, DC:monocyte-derived:Schuler_treatment: 0.41, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.41, DC:monocyte-derived:anti-DC-SIGN_2h: 0.4 |
| WK032-H13 | Macrophage:monocyte-derived:M-CSF/Pam3Cys | 0.20 | 214.77 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.54, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.52, DC:monocyte-derived:antiCD40/VAF347: 0.52, Macrophage:monocyte-derived:S._aureus: 0.51, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.5, DC:monocyte-derived:LPS: 0.5, DC:monocyte-derived:Galectin-1: 0.5, Monocyte:S._typhimurium_flagellin: 0.5, Macrophage:monocyte-derived:IFNa: 0.5, DC:monocyte-derived:CD40L: 0.49 |
| WK097-F4 | Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys | 0.18 | 214.22 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.45, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.45, Monocyte:S._typhimurium_flagellin: 0.44, DC:monocyte-derived:antiCD40/VAF347: 0.44, DC:monocyte-derived:Schuler_treatment: 0.43, Macrophage:monocyte-derived:S._aureus: 0.43, DC:monocyte-derived:LPS: 0.43, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.42, Monocyte:anti-FcgRIIB: 0.42, DC:monocyte-derived:CD40L: 0.42 |
| TM37-J11 | Monocyte:leukotriene_D4 | 0.11 | 212.27 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.42, DC:monocyte-derived:CD40L: 0.42, DC:monocyte-derived:mature: 0.41, DC:monocyte-derived:anti-DC-SIGN_2h: 0.41, Monocyte:leukotriene_D4: 0.41, Monocyte: 0.41, Macrophage:monocyte-derived:M-CSF/IFNg: 0.4, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.4, DC:monocyte-derived: 0.4, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.4 |
| WMK004-J5 | DC:monocyte-derived:Schuler_treatment | 0.14 | 208.90 | Raw ScoresDC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.38, DC:monocyte-derived:mature: 0.38, DC:monocyte-derived:CD40L: 0.37, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.37, DC:monocyte-derived:antiCD40/VAF347: 0.37, DC:monocyte-derived:anti-DC-SIGN_2h: 0.36, DC:monocyte-derived:Poly(IC): 0.36, DC:monocyte-derived:LPS: 0.36, DC:monocyte-derived:Galectin-1: 0.36 |
| WK014-J1 | Monocyte:S._typhimurium_flagellin | 0.19 | 208.18 | Raw ScoresMonocyte:S._typhimurium_flagellin: 0.41, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.4, DC:monocyte-derived:antiCD40/VAF347: 0.4, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.39, DC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.39, DC:monocyte-derived:LPS: 0.38, Macrophage:monocyte-derived:S._aureus: 0.38, DC:monocyte-derived:CD40L: 0.37, DC:monocyte-derived:Galectin-1: 0.37 |
| WK071-J13 | Monocyte:CD16+ | 0.10 | 208.14 | Raw ScoresMonocyte:leukotriene_D4: 0.38, Monocyte:CD16+: 0.38, Monocyte:CD16-: 0.38, Monocyte: 0.38, DC:monocyte-derived:anti-DC-SIGN_2h: 0.38, Monocyte:CD14+: 0.38, Monocyte:anti-FcgRIIB: 0.38, DC:monocyte-derived:mature: 0.37, Macrophage:monocyte-derived:M-CSF/IFNg: 0.37, DC:monocyte-derived:AM580: 0.37 |
| WK017-N2 | DC:monocyte-derived:LPS | 0.18 | 207.98 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.45, DC:monocyte-derived:antiCD40/VAF347: 0.45, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.44, DC:monocyte-derived:LPS: 0.44, Macrophage:monocyte-derived:S._aureus: 0.44, DC:monocyte-derived:Galectin-1: 0.43, DC:monocyte-derived:CD40L: 0.43, DC:monocyte-derived:Schuler_treatment: 0.43, DC:monocyte-derived:Poly(IC): 0.43, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.42 |
| WK073-D7 | Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys | 0.21 | 203.80 | Raw ScoresMacrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.47, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.47, Monocyte:S._typhimurium_flagellin: 0.46, DC:monocyte-derived:antiCD40/VAF347: 0.45, DC:monocyte-derived:LPS: 0.45, Monocyte:F._tularensis_novicida: 0.45, Macrophage:monocyte-derived:S._aureus: 0.44, DC:monocyte-derived:mature: 0.44, DC:monocyte-derived:Galectin-1: 0.44, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.44 |
| WK012-I16 | Macrophage:monocyte-derived:M-CSF/Pam3Cys | 0.23 | 201.56 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.49, Monocyte:S._typhimurium_flagellin: 0.49, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.48, DC:monocyte-derived:antiCD40/VAF347: 0.47, Monocyte:F._tularensis_novicida: 0.46, Macrophage:monocyte-derived:S._aureus: 0.45, Monocyte:anti-FcgRIIB: 0.45, DC:monocyte-derived:LPS: 0.45, DC:monocyte-derived:Galectin-1: 0.45, Macrophage:Alveolar:B._anthacis_spores: 0.45 |
| WK016-N17 | Macrophage:monocyte-derived:M-CSF | 0.09 | 201.33 | Raw ScoresMacrophage:monocyte-derived:M-CSF/Pam3Cys: 0.38, DC:monocyte-derived:antiCD40/VAF347: 0.38, DC:monocyte-derived:mature: 0.38, Macrophage:monocyte-derived:S._aureus: 0.38, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.38, DC:monocyte-derived:Schuler_treatment: 0.38, DC:monocyte-derived:LPS: 0.37, Macrophage:monocyte-derived: 0.37, Macrophage:monocyte-derived:M-CSF: 0.37, DC:monocyte-derived:anti-DC-SIGN_2h: 0.37 |
| WK039-M14 | DC:monocyte-derived:A._fumigatus_germ_tubes_6h | 0.19 | 192.82 | Raw ScoresDC:monocyte-derived:antiCD40/VAF347: 0.4, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.4, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.4, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.4, Monocyte:S._typhimurium_flagellin: 0.4, DC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:LPS: 0.39, DC:monocyte-derived:CD40L: 0.39, Macrophage:monocyte-derived:S._aureus: 0.38, DC:monocyte-derived:mature: 0.38 |
| WK066-D5 | DC:monocyte-derived:mature | 0.13 | 186.28 | Raw ScoresDC:monocyte-derived:mature: 0.4, DC:monocyte-derived:Schuler_treatment: 0.39, DC:monocyte-derived:CD40L: 0.37, DC:monocyte-derived:Poly(IC): 0.37, Monocyte:F._tularensis_novicida: 0.36, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.36, DC:monocyte-derived:immature: 0.36, DC:monocyte-derived:antiCD40/VAF347: 0.36, DC:monocyte-derived:LPS: 0.36, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.36 |
| WK099-G20 | Monocyte:S._typhimurium_flagellin | 0.16 | 185.58 | Raw ScoresMonocyte:S._typhimurium_flagellin: 0.39, Monocyte:F._tularensis_novicida: 0.39, Macrophage:monocyte-derived:M-CSF/Pam3Cys: 0.38, Macrophage:monocyte-derived:M-CSF/IFNg/Pam3Cys: 0.38, DC:monocyte-derived:antiCD40/VAF347: 0.37, DC:monocyte-derived:Schuler_treatment: 0.37, DC:monocyte-derived:A._fumigatus_germ_tubes_6h: 0.37, DC:monocyte-derived:LPS: 0.36, DC:monocyte-derived:CD40L: 0.36, Monocyte:anti-FcgRIIB: 0.36 |
| KK056-O12 | B_cell:immature | 0.09 | 183.02 | Raw ScoresPre-B_cell_CD34-: 0.49, Monocyte:leukotriene_D4: 0.49, Monocyte:CD16+: 0.49, Monocyte: 0.49, Monocyte:CD16-: 0.49, Monocyte:CD14+: 0.48, HSC_-G-CSF: 0.48, B_cell:immature: 0.48, DC:monocyte-derived:anti-DC-SIGN_2h: 0.48, Monocyte:anti-FcgRIIB: 0.48 |
Below shows the significant enrichments of this GEP for literature curated gene lists
This data was procured from existing single cell RNA-seq maps of neuroblastoma or related relevant data.
High ranks indicate this gene is a driver of this GEP.
These curated gene list are ranked by the P-value (on this GEP) of their constituent genes.
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
IFN Response (Kinker)
These marker genes were obtained in a pan cancer cell lines NMF analysis of scRNA-eq data in Kinker et al (PMID 33128048) - 10 metaprograms were recovered that manifest across cancers, this program contained interferon response genes.:
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.23e-03
Mean rank of genes in gene set: 1368
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| ISG15 | 0.0005842 | 714 | GTEx | DepMap | Descartes | 0.99 | 317.81 |
| IFIT2 | 0.0002893 | 1169 | GTEx | DepMap | Descartes | 0.30 | 20.71 |
| ISG20 | 0.0002402 | 1284 | GTEx | DepMap | Descartes | 0.79 | 35.51 |
| IFIT3 | 0.0001972 | 1408 | GTEx | DepMap | Descartes | 0.61 | 56.17 |
| IFIT1 | 0.0000357 | 2265 | GTEx | DepMap | Descartes | 0.05 | 2.40 |
p53 Dependent Senescence (Kinker)
These marker genes were obtained in a pan cancer cell lines NMF analysis of scRNA-eq data in Kinker et al (PMID 33128048) - 10 metaprograms were recovered that manifest across cancers.:
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.46e-02
Mean rank of genes in gene set: 689
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| NEAT1 | 0.0009244 | 445 | GTEx | DepMap | Descartes | 51.34 | 622.84 |
| CDKN1A | 0.0004124 | 933 | GTEx | DepMap | Descartes | 7.50 | 839.77 |
M-MDSC
These marker genes were curated for MDSC subtypes as reviewed in Veglia et. al. (PMID 33526920):
Wilcoxon ranksum test P-value for gene set overrepresentation: 2.10e-02
Mean rank of genes in gene set: 4373.73
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CD274 | 0.0013583 | 270 | GTEx | DepMap | Descartes | 0.71 | 49.16 |
| IL1B | 0.0010048 | 400 | GTEx | DepMap | Descartes | 72.94 | 11631.61 |
| VEGFA | 0.0007326 | 575 | GTEx | DepMap | Descartes | 3.79 | 82.58 |
| HIF1A | 0.0005969 | 696 | GTEx | DepMap | Descartes | 4.71 | 326.11 |
| TNFRSF10B | 0.0004683 | 843 | GTEx | DepMap | Descartes | 1.21 | 63.86 |
| IL10 | 0.0003590 | 1027 | GTEx | DepMap | Descartes | 5.89 | 708.93 |
| CD36 | -0.0000305 | 2994 | GTEx | DepMap | Descartes | 0.20 | 7.06 |
| TGFB1 | -0.0000467 | 3227 | GTEx | DepMap | Descartes | 1.13 | 92.43 |
| ARG1 | -0.0000614 | 3488 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| NOS2 | -0.0001091 | 4544 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ARG2 | -0.0001263 | 4951 | GTEx | DepMap | Descartes | 0.05 | 4.25 |
| CD84 | -0.0003231 | 9434 | GTEx | DepMap | Descartes | 1.60 | 50.96 |
| CD14 | -0.0003438 | 9781 | GTEx | DepMap | Descartes | 1.88 | 247.24 |
| STAT3 | -0.0004194 | 10827 | GTEx | DepMap | Descartes | 2.99 | 148.28 |
| TNF | -0.0028651 | 12549 | GTEx | DepMap | Descartes | 3.79 | 553.48 |
Below shows ranks on this GEP for literature curated gene lists for large gene sets
These include those reported as mesenchymal/adrenergic by Van Groningen et al.
High ranks indicate this gene is a driver of this GEP (note these results are not ordered).
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
VanGroningen Adrenergic Genes
Adrenergic marker genes from Supplementary Table 2 of Van Groningen et al. Nature Genetics 2017. These genes were identified by differential expression analysis of mesenchymal-like and adrenergic-like neuroblastoma cell lines.
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 8751.41
Median rank of genes in gene set: 10135
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| LYN | 0.0029673 | 82 | GTEx | DepMap | Descartes | 3.22 | 133.40 |
| DUSP4 | 0.0012352 | 307 | GTEx | DepMap | Descartes | 2.74 | 143.42 |
| TSPAN13 | 0.0011431 | 347 | GTEx | DepMap | Descartes | 0.42 | 59.41 |
| DAPK1 | 0.0010909 | 367 | GTEx | DepMap | Descartes | 0.81 | 33.44 |
| AP1S2 | 0.0009353 | 439 | GTEx | DepMap | Descartes | 1.65 | 104.14 |
| NET1 | 0.0009190 | 449 | GTEx | DepMap | Descartes | 0.73 | 30.86 |
| ST3GAL6 | 0.0005965 | 698 | GTEx | DepMap | Descartes | 0.93 | 60.60 |
| CXCR4 | 0.0005844 | 713 | GTEx | DepMap | Descartes | 6.35 | 807.93 |
| UCP2 | 0.0005705 | 727 | GTEx | DepMap | Descartes | 1.48 | 135.62 |
| NCS1 | 0.0005573 | 746 | GTEx | DepMap | Descartes | 0.53 | 26.33 |
| CKB | 0.0005114 | 795 | GTEx | DepMap | Descartes | 1.13 | 181.04 |
| ARL6IP1 | 0.0004851 | 823 | GTEx | DepMap | Descartes | 2.49 | 275.11 |
| GCH1 | 0.0004839 | 827 | GTEx | DepMap | Descartes | 1.49 | 121.27 |
| CHML | 0.0004451 | 882 | GTEx | DepMap | Descartes | 0.68 | 21.85 |
| SATB1 | 0.0004367 | 891 | GTEx | DepMap | Descartes | 1.70 | 56.08 |
| GNB1 | 0.0003929 | 971 | GTEx | DepMap | Descartes | 3.00 | 204.42 |
| HMGA1 | 0.0003546 | 1031 | GTEx | DepMap | Descartes | 1.85 | 163.63 |
| POLB | 0.0003198 | 1102 | GTEx | DepMap | Descartes | 0.38 | 71.70 |
| RAB33A | 0.0002511 | 1257 | GTEx | DepMap | Descartes | 0.20 | 43.97 |
| RBBP8 | 0.0002423 | 1279 | GTEx | DepMap | Descartes | 0.58 | 36.28 |
| CDC42EP3 | 0.0002287 | 1310 | GTEx | DepMap | Descartes | 2.34 | 111.69 |
| RUFY3 | 0.0001969 | 1410 | GTEx | DepMap | Descartes | 0.91 | 54.46 |
| CCNI | 0.0001760 | 1481 | GTEx | DepMap | Descartes | 7.43 | 595.50 |
| IGSF3 | 0.0001648 | 1526 | GTEx | DepMap | Descartes | 0.09 | 2.03 |
| FAM107B | 0.0001543 | 1568 | GTEx | DepMap | Descartes | 2.70 | 172.49 |
| RET | 0.0001350 | 1642 | GTEx | DepMap | Descartes | 0.03 | 2.26 |
| KLF7 | 0.0001269 | 1676 | GTEx | DepMap | Descartes | 0.61 | 16.15 |
| PPP2R3C | 0.0001211 | 1718 | GTEx | DepMap | Descartes | 0.35 | 38.17 |
| PBX3 | 0.0001143 | 1751 | GTEx | DepMap | Descartes | 0.56 | 42.07 |
| TIAM1 | 0.0001015 | 1815 | GTEx | DepMap | Descartes | 0.93 | 36.22 |
VanGroningen Mesenchymal Genes
Mesenchymal marker genes from Supplementary Table 2 of Van Groningen et al. Nature Genetics 2017. These genes were identified by differential expression analysis of mesenchymal-like and adrenergic-like neuroblastoma cell lines.
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 7485.99
Median rank of genes in gene set: 8978
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| OGFRL1 | 0.0057437 | 16 | GTEx | DepMap | Descartes | 3.32 | 95.67 |
| CMTM6 | 0.0054872 | 19 | GTEx | DepMap | Descartes | 6.99 | 565.46 |
| EDEM1 | 0.0039725 | 47 | GTEx | DepMap | Descartes | 2.36 | 98.40 |
| LITAF | 0.0036935 | 52 | GTEx | DepMap | Descartes | 20.35 | 2038.33 |
| INSIG1 | 0.0036552 | 54 | GTEx | DepMap | Descartes | 10.54 | 973.25 |
| ADAM19 | 0.0026029 | 100 | GTEx | DepMap | Descartes | 1.34 | 53.53 |
| GRN | 0.0025277 | 108 | GTEx | DepMap | Descartes | 8.30 | 894.95 |
| IL13RA1 | 0.0024343 | 113 | GTEx | DepMap | Descartes | 1.18 | 71.75 |
| PPT1 | 0.0020934 | 135 | GTEx | DepMap | Descartes | 2.99 | 153.38 |
| GPR137B | 0.0020519 | 140 | GTEx | DepMap | Descartes | 1.12 | 130.44 |
| SQSTM1 | 0.0019956 | 149 | GTEx | DepMap | Descartes | 11.40 | 960.15 |
| DSE | 0.0018527 | 170 | GTEx | DepMap | Descartes | 2.91 | 71.57 |
| PLSCR1 | 0.0015441 | 225 | GTEx | DepMap | Descartes | 1.80 | 229.01 |
| DUSP5 | 0.0015324 | 228 | GTEx | DepMap | Descartes | 3.38 | 408.53 |
| WNT5A | 0.0015041 | 236 | GTEx | DepMap | Descartes | 0.20 | 10.88 |
| NPC2 | 0.0013158 | 281 | GTEx | DepMap | Descartes | 4.18 | 708.63 |
| PTGER4 | 0.0013154 | 282 | GTEx | DepMap | Descartes | 2.59 | 184.20 |
| RAB31 | 0.0012175 | 317 | GTEx | DepMap | Descartes | 2.31 | 141.57 |
| MOB1A | 0.0011269 | 355 | GTEx | DepMap | Descartes | 2.13 | 100.19 |
| SH3BGRL | 0.0010693 | 380 | GTEx | DepMap | Descartes | 1.86 | 258.52 |
| ARPC1B | 0.0010336 | 390 | GTEx | DepMap | Descartes | 1.57 | 187.66 |
| RAP1B | 0.0010059 | 398 | GTEx | DepMap | Descartes | 1.84 | 33.36 |
| RGS10 | 0.0009725 | 416 | GTEx | DepMap | Descartes | 1.32 | 297.77 |
| TIMP1 | 0.0009414 | 436 | GTEx | DepMap | Descartes | 15.77 | 3906.93 |
| STK38L | 0.0009337 | 440 | GTEx | DepMap | Descartes | 0.51 | 23.44 |
| SEL1L3 | 0.0009209 | 447 | GTEx | DepMap | Descartes | 0.57 | 36.40 |
| ANXA5 | 0.0008848 | 467 | GTEx | DepMap | Descartes | 11.09 | 1584.69 |
| B2M | 0.0008523 | 485 | GTEx | DepMap | Descartes | 66.18 | 6605.76 |
| PRCP | 0.0007781 | 541 | GTEx | DepMap | Descartes | 1.14 | 33.46 |
| SDCBP | 0.0007539 | 557 | GTEx | DepMap | Descartes | 7.68 | 554.17 |
Descartes adrenocortical markers
Top 50 marker genes of adrenocortical cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/adrenocortical/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 9.93e-01
Mean rank of genes in gene set: 7742.61
Median rank of genes in gene set: 8635.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| PAPSS2 | 0.0004665 | 847 | GTEx | DepMap | Descartes | 0.60 | 34.96 |
| SCARB1 | 0.0000791 | 1942 | GTEx | DepMap | Descartes | 0.28 | 9.00 |
| SCAP | 0.0000197 | 2402 | GTEx | DepMap | Descartes | 0.17 | 9.13 |
| INHA | 0.0000079 | 2514 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SH3BP5 | -0.0000071 | 2684 | GTEx | DepMap | Descartes | 0.97 | 66.51 |
| FDXR | -0.0000499 | 3288 | GTEx | DepMap | Descartes | 0.03 | 2.09 |
| SLC1A2 | -0.0000561 | 3385 | GTEx | DepMap | Descartes | 0.09 | 1.48 |
| STAR | -0.0000729 | 3732 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SGCZ | -0.0000799 | 3886 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| BAIAP2L1 | -0.0001171 | 4733 | GTEx | DepMap | Descartes | 0.01 | 0.37 |
| FDX1 | -0.0001184 | 4760 | GTEx | DepMap | Descartes | 0.25 | 17.14 |
| FREM2 | -0.0001442 | 5402 | GTEx | DepMap | Descartes | 0.02 | 0.19 |
| PDE10A | -0.0001755 | 6142 | GTEx | DepMap | Descartes | 0.03 | 0.70 |
| TM7SF2 | -0.0001899 | 6478 | GTEx | DepMap | Descartes | 0.05 | 5.17 |
| HMGCS1 | -0.0002408 | 7665 | GTEx | DepMap | Descartes | 0.51 | 22.99 |
| GRAMD1B | -0.0002544 | 7986 | GTEx | DepMap | Descartes | 0.28 | 9.79 |
| APOC1 | -0.0002778 | 8499 | GTEx | DepMap | Descartes | 2.28 | 798.76 |
| CYB5B | -0.0002784 | 8514 | GTEx | DepMap | Descartes | 0.34 | 15.09 |
| JAKMIP2 | -0.0002900 | 8757 | GTEx | DepMap | Descartes | 0.08 | 0.98 |
| DHCR24 | -0.0002917 | 8794 | GTEx | DepMap | Descartes | 0.03 | 1.26 |
| POR | -0.0003125 | 9233 | GTEx | DepMap | Descartes | 0.58 | 56.78 |
| DNER | -0.0003147 | 9280 | GTEx | DepMap | Descartes | 0.01 | 1.05 |
| ERN1 | -0.0003365 | 9690 | GTEx | DepMap | Descartes | 0.64 | 19.55 |
| FRMD5 | -0.0003516 | 9902 | GTEx | DepMap | Descartes | 0.01 | 0.59 |
| SH3PXD2B | -0.0003901 | 10469 | GTEx | DepMap | Descartes | 0.15 | 6.10 |
| DHCR7 | -0.0004090 | 10717 | GTEx | DepMap | Descartes | 0.07 | 7.05 |
| GSTA4 | -0.0004671 | 11263 | GTEx | DepMap | Descartes | 0.07 | 5.41 |
| SLC16A9 | -0.0004774 | 11352 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| MSMO1 | -0.0004855 | 11405 | GTEx | DepMap | Descartes | 0.14 | 12.47 |
| PEG3 | -0.0004876 | 11422 | GTEx | DepMap | Descartes | 0.34 | NA |
Descartes chromaffin markers
Top 50 marker genes of chromaffin cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/chromaffin/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 8460.63
Median rank of genes in gene set: 9283
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| BASP1 | 0.0032520 | 66 | GTEx | DepMap | Descartes | 13.25 | 1873.30 |
| EYA4 | 0.0001330 | 1647 | GTEx | DepMap | Descartes | 0.03 | 1.79 |
| PLXNA4 | 0.0000387 | 2243 | GTEx | DepMap | Descartes | 0.01 | 0.21 |
| ANKFN1 | 0.0000072 | 2518 | GTEx | DepMap | Descartes | 0.02 | 1.24 |
| EPHA6 | 0.0000014 | 2587 | GTEx | DepMap | Descartes | 0.01 | 1.88 |
| HS3ST5 | -0.0000267 | 2937 | GTEx | DepMap | Descartes | 0.03 | 1.27 |
| GAL | -0.0001016 | 4388 | GTEx | DepMap | Descartes | 0.02 | 3.21 |
| KCNB2 | -0.0001195 | 4782 | GTEx | DepMap | Descartes | 0.01 | 1.01 |
| IL7 | -0.0001361 | 5190 | GTEx | DepMap | Descartes | 0.02 | 1.88 |
| RBFOX1 | -0.0001736 | 6098 | GTEx | DepMap | Descartes | 0.03 | 0.72 |
| SLC44A5 | -0.0001868 | 6400 | GTEx | DepMap | Descartes | 0.01 | 0.21 |
| TMEFF2 | -0.0001893 | 6468 | GTEx | DepMap | Descartes | 0.04 | 2.01 |
| PTCHD1 | -0.0002187 | 7136 | GTEx | DepMap | Descartes | 0.09 | 1.06 |
| RPH3A | -0.0002391 | 7624 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| RYR2 | -0.0002684 | 8282 | GTEx | DepMap | Descartes | 0.06 | 0.75 |
| NPY | -0.0002739 | 8396 | GTEx | DepMap | Descartes | 0.14 | 30.71 |
| GREM1 | -0.0002802 | 8541 | GTEx | DepMap | Descartes | 0.03 | 0.36 |
| MAB21L2 | -0.0002818 | 8581 | GTEx | DepMap | Descartes | 0.06 | 7.26 |
| EYA1 | -0.0002907 | 8769 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ALK | -0.0002995 | 8964 | GTEx | DepMap | Descartes | 0.02 | 0.33 |
| NTRK1 | -0.0003150 | 9283 | GTEx | DepMap | Descartes | 0.01 | 0.49 |
| TMEM132C | -0.0003225 | 9422 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| CNTFR | -0.0003279 | 9535 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| FAT3 | -0.0003414 | 9741 | GTEx | DepMap | Descartes | 0.01 | 0.08 |
| SYNPO2 | -0.0003672 | 10139 | GTEx | DepMap | Descartes | 0.22 | 3.16 |
| REEP1 | -0.0003828 | 10370 | GTEx | DepMap | Descartes | 0.06 | 3.58 |
| SLC6A2 | -0.0004371 | 10981 | GTEx | DepMap | Descartes | 0.02 | 0.66 |
| GAP43 | -0.0004391 | 11009 | GTEx | DepMap | Descartes | 0.41 | 36.50 |
| CNKSR2 | -0.0004398 | 11017 | GTEx | DepMap | Descartes | 0.01 | 0.14 |
| CCND1 | -0.0004997 | 11514 | GTEx | DepMap | Descartes | 0.97 | 51.68 |
Descartes Vascular_endothelial markers
Top 50 marker genes of Vascular_endothelial cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/vascular_endothelial/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 8.99e-01
Mean rank of genes in gene set: 7024.82
Median rank of genes in gene set: 7256.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CALCRL | 0.0012153 | 318 | GTEx | DepMap | Descartes | 0.68 | 23.34 |
| CRHBP | 0.0001725 | 1497 | GTEx | DepMap | Descartes | 0.04 | 4.28 |
| TMEM88 | 0.0000645 | 2040 | GTEx | DepMap | Descartes | 0.24 | 70.88 |
| NR5A2 | -0.0000800 | 3888 | GTEx | DepMap | Descartes | 0.01 | 0.28 |
| EHD3 | -0.0000804 | 3893 | GTEx | DepMap | Descartes | 0.01 | 0.35 |
| F8 | -0.0000870 | 4044 | GTEx | DepMap | Descartes | 0.06 | 1.51 |
| SHE | -0.0000940 | 4217 | GTEx | DepMap | Descartes | 0.06 | 1.27 |
| GALNT15 | -0.0000990 | 4329 | GTEx | DepMap | Descartes | 0.01 | NA |
| NPR1 | -0.0001446 | 5416 | GTEx | DepMap | Descartes | 0.01 | 0.15 |
| CEACAM1 | -0.0001466 | 5467 | GTEx | DepMap | Descartes | 0.03 | 1.37 |
| ROBO4 | -0.0001511 | 5590 | GTEx | DepMap | Descartes | 0.03 | 1.54 |
| IRX3 | -0.0001552 | 5666 | GTEx | DepMap | Descartes | 0.02 | 0.71 |
| RASIP1 | -0.0001634 | 5856 | GTEx | DepMap | Descartes | 0.04 | 4.22 |
| SLCO2A1 | -0.0001808 | 6266 | GTEx | DepMap | Descartes | 0.06 | 3.80 |
| CLDN5 | -0.0001812 | 6277 | GTEx | DepMap | Descartes | 0.01 | 0.82 |
| TEK | -0.0002022 | 6756 | GTEx | DepMap | Descartes | 0.01 | 0.05 |
| BTNL9 | -0.0002064 | 6845 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| PTPRB | -0.0002077 | 6881 | GTEx | DepMap | Descartes | 0.10 | 1.75 |
| MMRN2 | -0.0002155 | 7062 | GTEx | DepMap | Descartes | 0.02 | 1.28 |
| CHRM3 | -0.0002312 | 7451 | GTEx | DepMap | Descartes | 0.01 | 0.05 |
| MYRIP | -0.0002317 | 7462 | GTEx | DepMap | Descartes | 0.02 | 0.76 |
| NOTCH4 | -0.0002361 | 7554 | GTEx | DepMap | Descartes | 0.04 | 1.19 |
| ESM1 | -0.0002384 | 7606 | GTEx | DepMap | Descartes | 0.02 | 1.20 |
| KANK3 | -0.0002769 | 8481 | GTEx | DepMap | Descartes | 0.02 | 1.63 |
| SHANK3 | -0.0002788 | 8517 | GTEx | DepMap | Descartes | 0.07 | 1.24 |
| KDR | -0.0002810 | 8556 | GTEx | DepMap | Descartes | 0.03 | 0.54 |
| PLVAP | -0.0002974 | 8927 | GTEx | DepMap | Descartes | 0.17 | 11.50 |
| FLT4 | -0.0003011 | 9004 | GTEx | DepMap | Descartes | 0.01 | 0.22 |
| TIE1 | -0.0003015 | 9011 | GTEx | DepMap | Descartes | 0.07 | 3.88 |
| CDH5 | -0.0003056 | 9100 | GTEx | DepMap | Descartes | 0.02 | 0.72 |
Descartes stromal markers
Top 50 marker genes of stromal cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/stromal/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 9517.93
Median rank of genes in gene set: 9597
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| DKK2 | -0.0000291 | 2974 | GTEx | DepMap | Descartes | 0.01 | 0.69 |
| LAMC3 | -0.0000927 | 4185 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| GAS2 | -0.0000939 | 4214 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| RSPO3 | -0.0001724 | 6072 | GTEx | DepMap | Descartes | 0.02 | NA |
| SFRP2 | -0.0001943 | 6577 | GTEx | DepMap | Descartes | 0.06 | 9.25 |
| LRRC17 | -0.0002177 | 7113 | GTEx | DepMap | Descartes | 0.01 | 0.48 |
| HHIP | -0.0002202 | 7169 | GTEx | DepMap | Descartes | 0.02 | 0.63 |
| FREM1 | -0.0002205 | 7176 | GTEx | DepMap | Descartes | 0.01 | 0.17 |
| POSTN | -0.0002219 | 7217 | GTEx | DepMap | Descartes | 0.05 | 3.28 |
| CLDN11 | -0.0002259 | 7314 | GTEx | DepMap | Descartes | 0.02 | 1.17 |
| GLI2 | -0.0002472 | 7813 | GTEx | DepMap | Descartes | 0.01 | 0.38 |
| ITGA11 | -0.0002746 | 8419 | GTEx | DepMap | Descartes | 0.04 | 0.95 |
| ELN | -0.0002887 | 8716 | GTEx | DepMap | Descartes | 0.08 | 1.41 |
| ABCA6 | -0.0002999 | 8977 | GTEx | DepMap | Descartes | 0.07 | 1.96 |
| ACTA2 | -0.0003009 | 8996 | GTEx | DepMap | Descartes | 0.64 | 60.45 |
| PCDH18 | -0.0003011 | 9000 | GTEx | DepMap | Descartes | 0.03 | 0.69 |
| SCARA5 | -0.0003100 | 9184 | GTEx | DepMap | Descartes | 0.01 | 0.22 |
| ABCC9 | -0.0003122 | 9225 | GTEx | DepMap | Descartes | 0.02 | 0.51 |
| ADAMTSL3 | -0.0003194 | 9369 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| COL27A1 | -0.0003203 | 9380 | GTEx | DepMap | Descartes | 0.01 | 0.49 |
| PRICKLE1 | -0.0003240 | 9451 | GTEx | DepMap | Descartes | 0.03 | 1.32 |
| LOX | -0.0003317 | 9595 | GTEx | DepMap | Descartes | 0.03 | 0.93 |
| PAMR1 | -0.0003320 | 9599 | GTEx | DepMap | Descartes | 0.02 | 1.73 |
| EDNRA | -0.0003346 | 9656 | GTEx | DepMap | Descartes | 0.05 | 1.83 |
| CD248 | -0.0003426 | 9759 | GTEx | DepMap | Descartes | 0.03 | 1.53 |
| ADAMTS2 | -0.0003484 | 9851 | GTEx | DepMap | Descartes | 0.09 | 3.31 |
| IGFBP3 | -0.0003687 | 10159 | GTEx | DepMap | Descartes | 0.05 | 2.69 |
| COL1A1 | -0.0004044 | 10661 | GTEx | DepMap | Descartes | 5.58 | 251.61 |
| BICC1 | -0.0004361 | 10968 | GTEx | DepMap | Descartes | 0.03 | 0.89 |
| CCDC80 | -0.0004865 | 11411 | GTEx | DepMap | Descartes | 0.27 | 6.27 |
Descartes sympathoblasts markers
Top 50 marker genes of sympathoblasts cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/sympathoblasts/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 3.56e-01
Mean rank of genes in gene set: 6059
Median rank of genes in gene set: 5547.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| ST18 | 0.0013906 | 262 | GTEx | DepMap | Descartes | 0.20 | 6.48 |
| CCSER1 | 0.0013144 | 283 | GTEx | DepMap | Descartes | 0.11 | NA |
| GCH1 | 0.0004839 | 827 | GTEx | DepMap | Descartes | 1.49 | 121.27 |
| TIAM1 | 0.0001015 | 1815 | GTEx | DepMap | Descartes | 0.93 | 36.22 |
| SLC35F3 | 0.0000438 | 2199 | GTEx | DepMap | Descartes | 0.01 | 0.71 |
| EML6 | -0.0000054 | 2660 | GTEx | DepMap | Descartes | 0.05 | 0.55 |
| TBX20 | -0.0000084 | 2699 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SORCS3 | -0.0000403 | 3126 | GTEx | DepMap | Descartes | 0.01 | 0.40 |
| PACRG | -0.0000489 | 3270 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SPOCK3 | -0.0000652 | 3568 | GTEx | DepMap | Descartes | 0.01 | 0.68 |
| KSR2 | -0.0000911 | 4158 | GTEx | DepMap | Descartes | 0.01 | 0.11 |
| CDH18 | -0.0001036 | 4423 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| GALNTL6 | -0.0001149 | 4670 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| PENK | -0.0001187 | 4765 | GTEx | DepMap | Descartes | 0.01 | 1.45 |
| AGBL4 | -0.0001207 | 4809 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| GRM7 | -0.0001307 | 5067 | GTEx | DepMap | Descartes | 0.01 | 0.23 |
| PCSK2 | -0.0001421 | 5345 | GTEx | DepMap | Descartes | 0.02 | 0.76 |
| SLC24A2 | -0.0001477 | 5491 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| FGF14 | -0.0001490 | 5520 | GTEx | DepMap | Descartes | 0.01 | 0.07 |
| LAMA3 | -0.0001507 | 5575 | GTEx | DepMap | Descartes | 0.03 | 0.81 |
| CDH12 | -0.0001563 | 5690 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| CNTN3 | -0.0001630 | 5843 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| SLC18A1 | -0.0001826 | 6301 | GTEx | DepMap | Descartes | 0.01 | 0.45 |
| GRID2 | -0.0001840 | 6333 | GTEx | DepMap | Descartes | 0.01 | 0.28 |
| ROBO1 | -0.0002057 | 6834 | GTEx | DepMap | Descartes | 0.12 | 2.86 |
| KCTD16 | -0.0002135 | 7005 | GTEx | DepMap | Descartes | 0.03 | 0.25 |
| NTNG1 | -0.0002473 | 7817 | GTEx | DepMap | Descartes | 0.03 | 1.27 |
| TENM1 | -0.0002704 | 8320 | GTEx | DepMap | Descartes | 0.06 | NA |
| MGAT4C | -0.0002736 | 8386 | GTEx | DepMap | Descartes | 0.02 | 0.11 |
| UNC80 | -0.0003189 | 9352 | GTEx | DepMap | Descartes | 0.02 | 0.52 |
Descartes erythroblasts markers
Top 50 marker genes of erythroblasts cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/erythroblasts/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 4.63e-01
Mean rank of genes in gene set: 6212
Median rank of genes in gene set: 6309
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CAT | 0.0010691 | 381 | GTEx | DepMap | Descartes | 0.78 | 76.50 |
| DENND4A | 0.0010273 | 393 | GTEx | DepMap | Descartes | 1.87 | 53.80 |
| SPECC1 | 0.0006954 | 606 | GTEx | DepMap | Descartes | 0.25 | 5.35 |
| RAPGEF2 | 0.0006793 | 623 | GTEx | DepMap | Descartes | 2.11 | 71.16 |
| ABCB10 | 0.0000382 | 2246 | GTEx | DepMap | Descartes | 0.14 | 10.93 |
| RHD | 0.0000371 | 2255 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| TMCC2 | 0.0000204 | 2396 | GTEx | DepMap | Descartes | 0.07 | 4.86 |
| SLC25A21 | -0.0000367 | 3072 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| GCLC | -0.0000452 | 3207 | GTEx | DepMap | Descartes | 0.40 | 22.11 |
| ALAS2 | -0.0000865 | 4028 | GTEx | DepMap | Descartes | 0.01 | 0.52 |
| SLC4A1 | -0.0000868 | 4038 | GTEx | DepMap | Descartes | 0.01 | 0.09 |
| MARCH3 | -0.0001005 | 4363 | GTEx | DepMap | Descartes | 0.13 | NA |
| FECH | -0.0001061 | 4478 | GTEx | DepMap | Descartes | 0.07 | 1.67 |
| TFR2 | -0.0001163 | 4712 | GTEx | DepMap | Descartes | 0.01 | 1.26 |
| RGS6 | -0.0001829 | 6309 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| CPOX | -0.0001959 | 6611 | GTEx | DepMap | Descartes | 0.10 | 7.79 |
| SPTB | -0.0002034 | 6780 | GTEx | DepMap | Descartes | 0.01 | 0.20 |
| SOX6 | -0.0002816 | 8575 | GTEx | DepMap | Descartes | 0.01 | 0.36 |
| ANK1 | -0.0002900 | 8754 | GTEx | DepMap | Descartes | 0.03 | 0.37 |
| SNCA | -0.0003021 | 9035 | GTEx | DepMap | Descartes | 0.12 | 10.44 |
| BLVRB | -0.0003206 | 9385 | GTEx | DepMap | Descartes | 0.38 | 52.99 |
| MICAL2 | -0.0003260 | 9495 | GTEx | DepMap | Descartes | 0.13 | 3.36 |
| SELENBP1 | -0.0003266 | 9512 | GTEx | DepMap | Descartes | 0.01 | 1.24 |
| XPO7 | -0.0003415 | 9742 | GTEx | DepMap | Descartes | 0.18 | 10.29 |
| TRAK2 | -0.0004690 | 11278 | GTEx | DepMap | Descartes | 0.15 | 4.24 |
| EPB41 | -0.0005126 | 11598 | GTEx | DepMap | Descartes | 0.28 | 10.65 |
| TSPAN5 | -0.0005742 | 11935 | GTEx | DepMap | Descartes | 0.07 | 4.44 |
| GYPC | -0.0005871 | 11988 | GTEx | DepMap | Descartes | 0.83 | 88.37 |
| SLC25A37 | -0.0007468 | 12353 | GTEx | DepMap | Descartes | 0.86 | 36.65 |
| NA | NA | NA | NA | NA | NA | NA | NA |
Descartes myeloid markers
Top 50 marker genes of myeloid cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/myeloid/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.18e-01
Mean rank of genes in gene set: 5578.87
Median rank of genes in gene set: 3305
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CD74 | 0.0118294 | 1 | GTEx | DepMap | Descartes | 212.37 | 16090.36 |
| CST3 | 0.0068182 | 12 | GTEx | DepMap | Descartes | 23.92 | 1682.07 |
| CPVL | 0.0063634 | 14 | GTEx | DepMap | Descartes | 3.57 | 349.47 |
| HCK | 0.0031084 | 75 | GTEx | DepMap | Descartes | 2.97 | 268.22 |
| CTSS | 0.0019092 | 160 | GTEx | DepMap | Descartes | 6.21 | 380.98 |
| MARCH1 | 0.0016137 | 207 | GTEx | DepMap | Descartes | 0.66 | NA |
| FGL2 | 0.0015228 | 232 | GTEx | DepMap | Descartes | 3.91 | 198.95 |
| AXL | 0.0014546 | 250 | GTEx | DepMap | Descartes | 1.44 | 82.51 |
| PTPRE | 0.0012946 | 295 | GTEx | DepMap | Descartes | 1.94 | 69.79 |
| CYBB | 0.0011074 | 362 | GTEx | DepMap | Descartes | 2.27 | 120.51 |
| ATP8B4 | 0.0010000 | 402 | GTEx | DepMap | Descartes | 0.53 | 19.36 |
| CSF1R | 0.0009139 | 451 | GTEx | DepMap | Descartes | 1.94 | 105.20 |
| FGD2 | 0.0007960 | 522 | GTEx | DepMap | Descartes | 0.36 | 12.78 |
| TGFBI | 0.0004168 | 930 | GTEx | DepMap | Descartes | 3.54 | 194.06 |
| SLC9A9 | 0.0002540 | 1248 | GTEx | DepMap | Descartes | 0.31 | 19.80 |
| IFNGR1 | 0.0002479 | 1267 | GTEx | DepMap | Descartes | 1.00 | 89.71 |
| SFMBT2 | 0.0002424 | 1278 | GTEx | DepMap | Descartes | 0.55 | 11.42 |
| ADAP2 | 0.0000878 | 1884 | GTEx | DepMap | Descartes | 0.38 | 31.11 |
| MSR1 | -0.0000493 | 3277 | GTEx | DepMap | Descartes | 0.87 | 66.07 |
| RGL1 | -0.0000529 | 3333 | GTEx | DepMap | Descartes | 0.55 | 27.33 |
| RBPJ | -0.0001410 | 5306 | GTEx | DepMap | Descartes | 1.92 | 71.90 |
| ABCA1 | -0.0002750 | 8427 | GTEx | DepMap | Descartes | 2.01 | 46.03 |
| CD14 | -0.0003438 | 9781 | GTEx | DepMap | Descartes | 1.88 | 247.24 |
| MERTK | -0.0003486 | 9857 | GTEx | DepMap | Descartes | 0.11 | 6.10 |
| FMN1 | -0.0003531 | 9933 | GTEx | DepMap | Descartes | 0.21 | 4.11 |
| HRH1 | -0.0004016 | 10613 | GTEx | DepMap | Descartes | 0.06 | 2.23 |
| SPP1 | -0.0004100 | 10728 | GTEx | DepMap | Descartes | 0.38 | 59.31 |
| CTSB | -0.0004177 | 10817 | GTEx | DepMap | Descartes | 10.56 | 655.95 |
| ITPR2 | -0.0004381 | 10995 | GTEx | DepMap | Descartes | 0.51 | 7.97 |
| CTSC | -0.0004998 | 11515 | GTEx | DepMap | Descartes | 1.57 | 52.09 |
Descartes Schwann markers
Top 50 marker genes of Schwann cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/schwann/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 8405.11
Median rank of genes in gene set: 9122.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| MARCKS | 0.0022767 | 121 | GTEx | DepMap | Descartes | 14.87 | 897.57 |
| VIM | 0.0002722 | 1208 | GTEx | DepMap | Descartes | 28.59 | 2759.52 |
| IL1RAPL2 | -0.0000254 | 2919 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| TRPM3 | -0.0000394 | 3115 | GTEx | DepMap | Descartes | 0.02 | 0.52 |
| GAS7 | -0.0000561 | 3386 | GTEx | DepMap | Descartes | 0.69 | 20.11 |
| HMGA2 | -0.0000973 | 4283 | GTEx | DepMap | Descartes | 0.01 | 0.44 |
| MDGA2 | -0.0001072 | 4503 | GTEx | DepMap | Descartes | 0.01 | 0.42 |
| ERBB4 | -0.0001100 | 4564 | GTEx | DepMap | Descartes | 0.01 | 0.14 |
| LRRTM4 | -0.0001239 | 4887 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| IL1RAPL1 | -0.0001500 | 5555 | GTEx | DepMap | Descartes | 0.01 | 0.14 |
| NRXN3 | -0.0001673 | 5944 | GTEx | DepMap | Descartes | 0.01 | 0.18 |
| MPZ | -0.0001975 | 6648 | GTEx | DepMap | Descartes | 0.03 | 2.69 |
| XKR4 | -0.0002016 | 6743 | GTEx | DepMap | Descartes | 0.03 | 0.31 |
| SLC35F1 | -0.0002074 | 6865 | GTEx | DepMap | Descartes | 0.02 | 0.96 |
| SOX5 | -0.0002291 | 7404 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| PLCE1 | -0.0002546 | 7990 | GTEx | DepMap | Descartes | 0.01 | 0.04 |
| PTN | -0.0002565 | 8029 | GTEx | DepMap | Descartes | 0.03 | 4.16 |
| ADAMTS5 | -0.0002583 | 8071 | GTEx | DepMap | Descartes | 0.01 | 0.44 |
| SCN7A | -0.0002599 | 8103 | GTEx | DepMap | Descartes | 0.03 | 1.00 |
| GRIK3 | -0.0002929 | 8821 | GTEx | DepMap | Descartes | 0.01 | 0.26 |
| COL25A1 | -0.0003017 | 9024 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ERBB3 | -0.0003060 | 9104 | GTEx | DepMap | Descartes | 0.02 | 0.86 |
| EGFLAM | -0.0003077 | 9141 | GTEx | DepMap | Descartes | 0.05 | 0.94 |
| PTPRZ1 | -0.0003187 | 9349 | GTEx | DepMap | Descartes | 0.01 | 0.16 |
| PLP1 | -0.0003265 | 9508 | GTEx | DepMap | Descartes | 0.04 | 2.88 |
| OLFML2A | -0.0003301 | 9568 | GTEx | DepMap | Descartes | 0.03 | 0.58 |
| FIGN | -0.0003483 | 9845 | GTEx | DepMap | Descartes | 0.03 | 0.60 |
| GFRA3 | -0.0003495 | 9875 | GTEx | DepMap | Descartes | 0.05 | 4.23 |
| NRXN1 | -0.0003849 | 10397 | GTEx | DepMap | Descartes | 0.02 | 0.83 |
| SORCS1 | -0.0003896 | 10459 | GTEx | DepMap | Descartes | 0.03 | 0.85 |
Descartes Megakaryocytes markers
Top 50 marker genes of Megakaryocytes cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/megakaryocytes/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 3.22e-02
Mean rank of genes in gene set: 5278.36
Median rank of genes in gene set: 5516
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| ACTB | 0.0036115 | 58 | GTEx | DepMap | Descartes | 43.66 | 3929.12 |
| TMSB4X | 0.0023233 | 117 | GTEx | DepMap | Descartes | 41.35 | 5363.29 |
| PLEK | 0.0022423 | 124 | GTEx | DepMap | Descartes | 9.03 | 847.02 |
| PSTPIP2 | 0.0014360 | 254 | GTEx | DepMap | Descartes | 0.94 | 67.56 |
| RAP1B | 0.0010059 | 398 | GTEx | DepMap | Descartes | 1.84 | 33.36 |
| FLNA | 0.0007195 | 585 | GTEx | DepMap | Descartes | 5.42 | 155.81 |
| UBASH3B | 0.0007023 | 600 | GTEx | DepMap | Descartes | 0.56 | 15.15 |
| LIMS1 | 0.0003991 | 952 | GTEx | DepMap | Descartes | 3.38 | 194.64 |
| FERMT3 | 0.0003847 | 990 | GTEx | DepMap | Descartes | 1.03 | 89.01 |
| P2RX1 | 0.0003287 | 1085 | GTEx | DepMap | Descartes | 0.13 | 15.06 |
| MCTP1 | 0.0003133 | 1119 | GTEx | DepMap | Descartes | 0.66 | 26.44 |
| HIPK2 | 0.0002456 | 1269 | GTEx | DepMap | Descartes | 0.92 | 11.06 |
| CD9 | 0.0001656 | 1524 | GTEx | DepMap | Descartes | 0.87 | 122.31 |
| ZYX | 0.0001400 | 1623 | GTEx | DepMap | Descartes | 2.62 | 248.06 |
| PRKAR2B | 0.0000283 | 2329 | GTEx | DepMap | Descartes | 0.24 | 11.62 |
| TUBB1 | 0.0000233 | 2363 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| THBS1 | -0.0000102 | 2719 | GTEx | DepMap | Descartes | 13.39 | 480.72 |
| TGFB1 | -0.0000467 | 3227 | GTEx | DepMap | Descartes | 1.13 | 92.43 |
| ITGB3 | -0.0000595 | 3454 | GTEx | DepMap | Descartes | 0.01 | 0.57 |
| SLC2A3 | -0.0000823 | 3940 | GTEx | DepMap | Descartes | 2.86 | 159.51 |
| TLN1 | -0.0001025 | 4402 | GTEx | DepMap | Descartes | 2.19 | 54.50 |
| TRPC6 | -0.0001047 | 4450 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| MED12L | -0.0001487 | 5516 | GTEx | DepMap | Descartes | 0.02 | 0.75 |
| GP1BA | -0.0001638 | 5865 | GTEx | DepMap | Descartes | 0.02 | 1.60 |
| SPN | -0.0001761 | 6157 | GTEx | DepMap | Descartes | 0.51 | 17.70 |
| RAB27B | -0.0001801 | 6252 | GTEx | DepMap | Descartes | 0.01 | 0.33 |
| SLC24A3 | -0.0001883 | 6438 | GTEx | DepMap | Descartes | 0.00 | 0.00 |
| ARHGAP6 | -0.0002055 | 6830 | GTEx | DepMap | Descartes | 0.01 | 0.16 |
| ITGA2B | -0.0002307 | 7438 | GTEx | DepMap | Descartes | 0.01 | 0.19 |
| STON2 | -0.0002448 | 7761 | GTEx | DepMap | Descartes | 0.09 | 3.04 |
Descartes Lyphoid markers
Top 50 marker genes of Lyphoid cells in the Decartes fetal adrenal single cell map (https://atlas.brotmanbaty.org/bbi/human-gene-expression-during-development/cell/lymphoid/in/adrenal)
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.00e+00
Mean rank of genes in gene set: 9162.5
Median rank of genes in gene set: 11324.5
Rank on gene expression program of top 30 genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Descartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| TMSB10 | 0.0024843 | 110 | GTEx | DepMap | Descartes | 19.56 | 10246.01 |
| LCP1 | 0.0020333 | 143 | GTEx | DepMap | Descartes | 8.65 | 429.27 |
| B2M | 0.0008523 | 485 | GTEx | DepMap | Descartes | 66.18 | 6605.76 |
| BCL2 | 0.0007522 | 558 | GTEx | DepMap | Descartes | 1.22 | 35.75 |
| CD44 | 0.0007312 | 578 | GTEx | DepMap | Descartes | 16.10 | 750.28 |
| MSN | 0.0001014 | 1816 | GTEx | DepMap | Descartes | 2.76 | 158.16 |
| SORL1 | 0.0000114 | 2479 | GTEx | DepMap | Descartes | 1.87 | 40.29 |
| DOCK10 | -0.0001144 | 4662 | GTEx | DepMap | Descartes | 0.75 | 18.24 |
| CELF2 | -0.0001350 | 5164 | GTEx | DepMap | Descartes | 1.55 | 42.24 |
| FOXP1 | -0.0001363 | 5196 | GTEx | DepMap | Descartes | 2.13 | 59.21 |
| PLEKHA2 | -0.0002017 | 6746 | GTEx | DepMap | Descartes | 1.21 | 48.91 |
| RAP1GAP2 | -0.0002106 | 6935 | GTEx | DepMap | Descartes | 0.14 | 5.58 |
| NCALD | -0.0002875 | 8694 | GTEx | DepMap | Descartes | 0.06 | 3.50 |
| ARHGAP15 | -0.0003513 | 9897 | GTEx | DepMap | Descartes | 0.30 | 26.32 |
| ANKRD44 | -0.0003662 | 10123 | GTEx | DepMap | Descartes | 0.31 | 7.23 |
| MBNL1 | -0.0003886 | 10446 | GTEx | DepMap | Descartes | 2.13 | 66.66 |
| RCSD1 | -0.0003888 | 10448 | GTEx | DepMap | Descartes | 0.54 | 18.72 |
| BACH2 | -0.0004006 | 10599 | GTEx | DepMap | Descartes | 0.10 | 2.82 |
| IKZF1 | -0.0004017 | 10615 | GTEx | DepMap | Descartes | 0.81 | 24.22 |
| MCTP2 | -0.0004656 | 11253 | GTEx | DepMap | Descartes | 0.02 | 0.08 |
| SCML4 | -0.0004718 | 11309 | GTEx | DepMap | Descartes | 0.05 | 0.74 |
| TOX | -0.0004759 | 11340 | GTEx | DepMap | Descartes | 0.06 | 2.52 |
| CCND3 | -0.0005081 | 11570 | GTEx | DepMap | Descartes | 0.26 | 16.37 |
| SKAP1 | -0.0005266 | 11682 | GTEx | DepMap | Descartes | 0.02 | 2.30 |
| SAMD3 | -0.0005417 | 11782 | GTEx | DepMap | Descartes | 0.05 | 1.26 |
| SP100 | -0.0005428 | 11795 | GTEx | DepMap | Descartes | 0.87 | 32.40 |
| STK39 | -0.0005944 | 12013 | GTEx | DepMap | Descartes | 0.16 | 5.26 |
| ITPKB | -0.0006171 | 12076 | GTEx | DepMap | Descartes | 0.84 | 33.43 |
| PITPNC1 | -0.0007222 | 12310 | GTEx | DepMap | Descartes | 0.41 | 11.97 |
| ABLIM1 | -0.0007515 | 12359 | GTEx | DepMap | Descartes | 0.24 | 4.54 |
Below shows the ranks on this GEP for immune marker gene lists curated by CellTypist (https://www.celltypist.org/encyclopedia).
High ranks indicate this gene is a driver of this GEP.
These curated gene list are ranked by P-value (on this GEP) of their constituent genes.
The Mean Count column shows the mean read count in cells scoring highly (H > 50) on this gene expression program.
Monocytes: Monocytes (model markers)
myeloid mononuclear recirculating leukocytes that are capable of differentiating into macrophages and myeloid lineage dendritic cells:
Wilcoxon ranksum test P-value for gene set overrepresentation: 1.79e-03
Mean rank of genes in gene set: 183
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| SAT1 | 0.0043208 | 37 | GTEx | DepMap | Descartes | 32.39 | 7411.72 |
| TYROBP | 0.0032442 | 67 | GTEx | DepMap | Descartes | 11.88 | 4934.36 |
| NEAT1 | 0.0009244 | 445 | GTEx | DepMap | Descartes | 51.34 | 622.84 |
DC: Migratory DCs (curated markers)
migratory dendritic cells that transport antigens to the draining lymph nodes during both homeostatic conditions and infections:
Wilcoxon ranksum test P-value for gene set overrepresentation: 7.31e-03
Mean rank of genes in gene set: 21
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CCR7 | 0.0102872 | 4 | GTEx | DepMap | Descartes | 17.23 | 2204.68 |
| EBI3 | 0.0042778 | 38 | GTEx | DepMap | Descartes | 1.11 | 215.29 |
Monocytes: Classical monocytes (model markers)
CD14+ myeloid mononuclear recirculating leukocytes that are capable of differentiating into macrophages and myeloid lineage dendritic cells:
Wilcoxon ranksum test P-value for gene set overrepresentation: 7.55e-03
Mean rank of genes in gene set: 1192.67
Rank on gene expression program of genes in gene set:
| Genes | Weight | Rank | GTEx | DepMap | Decartes | Mean.Counts | Mean.TPM |
|---|---|---|---|---|---|---|---|
| CD207 | 0.0042092 | 39 | GTEx | DepMap | Descartes | 0.27 | 32.62 |
| CST7 | 0.0008514 | 486 | GTEx | DepMap | Descartes | 1.17 | 308.89 |
| EEF1A1 | -0.0000356 | 3053 | GTEx | DepMap | Descartes | 15.13 | 519.06 |